Tirandamycin
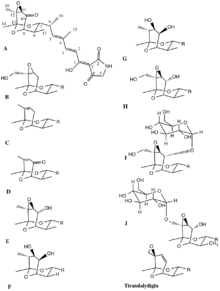
The tirandamycins are a small group of natural products that contain a bicyclic ketal system and a tetramic acid moiety, the latter of which is found in different natural products from a variety of sources and which is characterized by a 2,4-pyrrolidinedione ring system.[1] Members of this structural family have shown a wide range of biological activities like in antiparasitic, antifungal and anti-HIV evaluations, and furthermore, have shown potential usefulness because of their potent antibacterial properties.[2] Streptolydigin, an analogue of the tirandamycins, is known to function as an antibacterial agent through inhibiting the chain initiation and elongation steps RNA polymerase transcription.[3] The structural diversity in the tirandamycin family originates from the different oxidation patterns observed in the bicycic ketal system, and these modifications are determinant features for the bioactivity associated with these molecules.[4][5]
Biosynthesis
In the first study that looked at the gene cluster for tirandamycin production, Carlson et al. used primers specific for ketosynthase (KS) domains and CYP450 enzymes to probe the DNA of Streptomyces sp. 307-9, a previously determined producer of various tirandamycin analogues.[6] They found that the tirandamycin gene cluster is a PKS-NRPS hybrid that codes for three proteins with two, two and four PKS modules, and one other protein containing an NRPS module. Also, that the modules 0, 2, 6 and 7 AT domains are specific for loading or extending with malonate, while modules 1, 3, 4 and 5 are specific for methyl-malonate. The A domain in the NRPS module is specific for the amino acid glycine (See Figure 2). The cyclizations to form the tetramic acids’ 2,4-pyrrolidinedione ring and the bicyclic ketal system, as well as the oxidative transformations in the bicyclic skeleton, were suggested but further experimental evidence was needed.
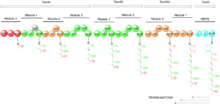
In another study Mo et al. characterized the biosynthetic gene cluster of the tirandamycins in Streptomyces sp. SCSIO1666 and described the function of one of the encoded proteins as a flavin-dependent oxidoreductase.[7] This enzyme was shown to be responsible for an oxidative transformation step (namely the 10-hydroxy dehydrogenation) to an intermediate that eventually leads to the ultimate product of the pathway, suggested to be tirandamycin B. Through a study of the metabolites produced after gene inactivation of the flavin-dependent enzyme and an in vitro characterization of the activity of the enzyme, they were able to conclude that the enzyme oxidizes tirandamycin E or F to tirandamycin A or D (See Figure 3).

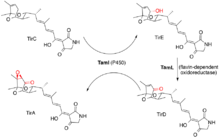
In the same year, Carlson et al. published another article that clarified even more the mechanisms involved in the generation of the oxidized metabolites.[8] They studied the action of a P450 enzyme contained in the pathway (TamI) by purifying it from a recombinant host, and saw, in vitro, that it can oxidize multiple oxidation steps, namely tirandamycin A into tirandamycin B and tirandamycin D into tirandamycin A, which correspond to two hydroxylations and one epoxidation reaction. This was the first time the versatile action of a single P450 enzyme was reported. The authors also evaluated the in vitro action of the flavin-dependent oxidoreductase, formerly characterized by Mo et al., against intermediates alone and with the presence of the TamI P450, and were able to show that these enzymes work together: TamI first hydroxylates the C10 of tirandamycin C to form tirandamycin E, then the flavin-dependent enzyme converts further oxidizes C10 into a carbonyl to form tirandamycin D, which then becomes a substrate for the TamI P450 that inserts an epoxide in the C11/C12 olefin (see Figure 4).
External links
References
- ↑ Xuhua Mo, Qinglian Lib and Jianhua Ju. Naturally occurring tetramic acid products: isolation, structure elucidation and biological activity. RSC Adv., 2014, 4, 50566-50593
- ↑ Jacob C. Carlson, Shengying Li, Douglas A. Burr, and David H. Sherman. Isolation and Characterization of Tirandamycins from a Marine-Derived Streptomyces sp. J. Nat. Prod. 2009, 72, 2076–2079
- ↑ Dmitry Temiakov, Nikolay Zenkin, Marina N. Vassylyeva, Anna Perederina, Tahir H. Tahirov, Ekaterina Kashkina, Maria Savkina, Savva Zorov, Vadim Nikiforov, Noriyuki Igarashi, Naohiro Matsugaki, Soichi Wakatsuki, Konstantin Severinov, and Dmitry G. Vassylyev. Structural Basis of Transcription Inhibition by Antibiotic Streptolydigin. Mol. Cell, 2009, 19, 655-666
- ↑ Dmitry Temiakov, Nikolay Zenkin, Marina N. Vassylyeva, Anna Perederina, Tahir H. Tahirov, Ekaterina Kashkina, Maria Savkina, Savva Zorov, Vadim Nikiforov, Noriyuki Igarashi, Naohiro Matsugaki, Soichi Wakatsuki, Konstantin Severinov, and Dmitry G. Vassylyev. Structural Basis of Transcription Inhibition by Antibiotic Streptolydigin. Mol. Cell, 2009, 19, 655-666
- ↑ Jacob C. Carlson, J. L. Fortman, Yojiro Anzai, Shengying Li, Douglas A. Burr, and David H. Sherman. Identification of the Tirandamycin Biosynthetic Gene Cluster from Streptomyces sp. 307-9 . ChemBioChem 2010, 11, 564 – 572
- ↑ Jacob C. Carlson, J. L. Fortman, Yojiro Anzai, Shengying Li, Douglas A. Burr, and David H. Sherman. Identification of the Tirandamycin Biosynthetic Gene Cluster from Streptomyces sp. 307-9 . ChemBioChem 2010, 11, 564 – 572
- ↑ Xuhua Mo, Hongbo Huang, Junying Ma, Zhongwen Wang, Bo Wang, Si Zhang, Changsheng Zhang, and Jianhua Ju. Characterization of TrdL as a 10-Hydroxy Dehydrogenase and Generation of New Analogues from a Tirandamycin Biosynthetic Pathway. Org. Lett. 2011, 13, 2212-2215
- ↑ Jacob C. Carlson, Shengying Li, Shamila S. Gunatilleke, Yojiro Anzai1, Douglas A. Burr, Larissa M. Podust, and David H. Sherman. Tirandamycin biosynthesis is mediated by co-dependent oxidative enzymes. Nat. Chem. 2011, 3, 628−633