ECHS1
| ECHS1 | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 |
|||||||||||||||||
| |||||||||||||||||
| Identifiers | |||||||||||||||||
| Aliases | ECHS1, SCEH, ECHS1D, enoyl-CoA hydratase, short chain, 1, mitochondrial | ||||||||||||||||
| External IDs | MGI: 2136460 HomoloGene: 3018 GeneCards: ECHS1 | ||||||||||||||||
| |||||||||||||||||
| RNA expression pattern | |||||||||||||||||
 | |||||||||||||||||
| More reference expression data | |||||||||||||||||
| Orthologs | |||||||||||||||||
| Species | Human | Mouse | |||||||||||||||
| Entrez | |||||||||||||||||
| Ensembl | |||||||||||||||||
| UniProt | |||||||||||||||||
| RefSeq (mRNA) | |||||||||||||||||
| RefSeq (protein) | |||||||||||||||||
| Location (UCSC) | Chr 10: 133.36 – 133.37 Mb | Chr 7: 140.11 – 140.12 Mb | |||||||||||||||
| PubMed search | [1] | [2] | |||||||||||||||
| Wikidata | |||||||||||||||||
| View/Edit Human | View/Edit Mouse |
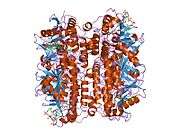
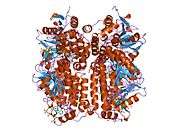
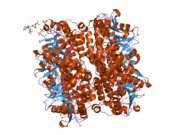
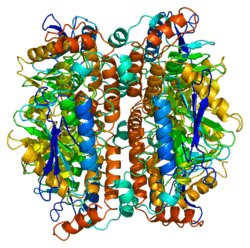
Enoyl Coenzyme A hydratase, short chain, 1, mitochondrial, also known as ECHS1, is a human gene.[3]
The protein encoded by this gene functions in the second step of the mitochondrial fatty acid beta-oxidation pathway. It catalyzes the hydration of 2-trans-enoyl-coenzyme A (CoA) intermediates to L-3-hydroxyacyl-CoAs. The gene product is a member of the hydratase/isomerase superfamily. It localizes to the mitochondrial matrix. Transcript variants utilizing alternative transcription initiation sites have been described in the literature.[3]
Structure
The ECHS1 gene is approximately 11 kb in length, and is composed of eight exons, with exons I and VIII containing the 5'- and 3'-untranslated regions, respectively. There are two major transcription start sites, located 62 and 63 bp upstream of the translation codon, were mapped by primer extension analysis. The 5'-flanking region of the ECHS1 gene is GC-rich and contains several copies of the SP1 binding motive but no typical TATA or CAAT boxes are apparent. Alu repeat elements have been identified within the region -1052/-770 relative to the cap site and in intron 7.[4] The precursor polypeptide contains 290 amino acid residues, with an N-terminal presequence of 29 residues, a 5'-untranslated sequence of 21 bp and a 3'-untranslated sequence of 391 bp.[5]
Function
Enoyl-CoA hydratase (ECH) catalyzes the second step in beta-oxidation pathway of fatty acid metabolism. The enzyme is involved in the formation of a β-hydroxyacyl-CoA thioester. The two catalytic glutamic acid residues are believed to act in concert to activate a water molecule, while Gly-141 is proposed to be involved in substrate activation. There are two potent inhibitors of ECHS, which irreversibly inactivate the enzyme via covalent adduct formation.[6]
Clinical significance
Enoyl-CoA hydratase short chain has been confirmed to interact with STAT3, such that ECHS1 specifically represses STAT3 activity by inhibiting STAT3 phosphorylation.[7] STAT3 can act as both an oncogene and a tumor suppressor. ECHS1 itself has shown to occur in many cancers, particularly in hypatocellular carcinoma (HCC) development;[8] both exogenous and endogenous forms of ECHS1 bind to HBs and induce apoptosis as a result. This means that ECHS1 may be used in the future as a therapy for patients with HBV-related hepatitis or HCC.[9]
References
- ↑ "Human PubMed Reference:".
- ↑ "Mouse PubMed Reference:".
- 1 2 "Entrez Gene: ECHS1 enoyl Coenzyme A hydratase, short chain, 1, mitochondrial".
- ↑ Janssen, U; Davis, E. M.; Le Beau, M. M.; Stoffel, W (1997). "Human mitochondrial enoyl-CoA hydratase gene (ECHS1): Structural organization and assignment to chromosome 10q26.2-q26.3". Genomics. 40 (3): 470–5. doi:10.1006/geno.1996.4597. PMID 9073515.
- ↑ Kanazawa, M; Ohtake, A; Abe, H; Yamamoto, S; Satoh, Y; Takayanagi, M; Niimi, H; Mori, M; Hashimoto, T (1993). "Molecular cloning and sequence analysis of the cDNA for human mitochondrial short-chain enoyl-CoA hydratase". Enzyme & protein. 47 (1): 9–13. PMID 8012501.
- ↑ Agnihotri, G; Liu, H. W. (2003). "Enoyl-CoA hydratase. Reaction, mechanism, and inhibition". Bioorganic & Medicinal Chemistry. 11 (1): 9–20. doi:10.1016/s0968-0896(02)00333-4. PMID 12467702.
- ↑ Chang, Y; Wang, S. X.; Wang, Y. B.; Zhou, J; Li, W. H.; Wang, N; Fang, D. F.; Li, H. Y.; Li, A. L.; Zhang, X. M.; Zhang, W. N. (2013). "ECHS1 interacts with STAT3 and negatively regulates STAT3 signaling". FEBS Letters. 587 (6): 607–13. doi:10.1016/j.febslet.2013.02.005. PMID 23416296.
- ↑ Zhu, X. S.; Dai, Y. C.; Chen, Z. X.; Xie, J. P.; Zeng, W; Lin, Y. Y.; Tan, Q. H. (2013). "Knockdown of ECHS1 protein expression inhibits hepatocellular carcinoma cell proliferation via suppression of Akt activity". Critical reviews in eukaryotic gene expression. 23 (3): 275–82. doi:10.1615/critreveukaryotgeneexpr.2013007531. PMID 23879543.
- ↑ Xiao, C. X.; Yang, X. N.; Huang, Q. W.; Zhang, Y. Q.; Lin, B. Y.; Liu, J. J.; Liu, Y. P.; Jazag, A; Guleng, B; Ren, J. L. (2013). "ECHS1 acts as a novel HBs Ag-binding protein enhancing apoptosis through the mitochondrial pathway in HepG2 cells". Cancer Letters. 330 (1): 67–73. doi:10.1016/j.canlet.2012.11.030. PMID 23178449.
Further reading
- Hochstrasser DF, Frutiger S, Paquet N, et al. (1993). "Human liver protein map: a reference database established by microsequencing and gel comparison.". Electrophoresis. 13 (12): 992–1001. doi:10.1002/elps.11501301201. PMID 1286669.
- Dawson SJ, White LA (1992). "Treatment of Haemophilus aphrophilus endocarditis with ciprofloxacin.". J. Infect. 24 (3): 317–20. doi:10.1016/S0163-4453(05)80037-4. PMID 1602151.
- Li J, Norwood DL, Mao LF, Schulz H (1991). "Mitochondrial metabolism of valproic acid.". Biochemistry. 30 (2): 388–94. doi:10.1021/bi00216a012. PMID 1988037.
- Jackson S, Schaefer J, Middleton B, Turnbull DM (1995). "Characterisation of a novel enzyme of human fatty acid beta-oxidation: a matrix-associated, mitochondrial 2-enoyl-CoA hydratase.". Biochem. Biophys. Res. Commun. 214 (1): 247–53. doi:10.1006/bbrc.1995.2281. PMID 7669045.
- Kanazawa M, Ohtake A, Abe H, et al. (1994). "Molecular cloning and sequence analysis of the cDNA for human mitochondrial short-chain enoyl-CoA hydratase.". Enzyme Protein. 47 (1): 9–13. PMID 8012501.
- Janssen U, Davis EM, Le Beau MM, Stoffel W (1997). "Human mitochondrial enoyl-CoA hydratase gene (ECHS1): structural organization and assignment to chromosome 10q26.2-q26.3.". Genomics. 40 (3): 470–5. doi:10.1006/geno.1996.4597. PMID 9073515.
- Hubbard MJ, McHugh NJ (2001). "Human ERp29: isolation, primary structural characterisation and two-dimensional gel mapping.". Electrophoresis. 21 (17): 3785–96. doi:10.1002/1522-2683(200011)21:17<3785::AID-ELPS3785>3.0.CO;2-2. PMID 11271497.
- Jiang LQ, Wen SJ, Wang HY, Chen LY (2003). "Screening the proteins that interact with calpain in a human heart cDNA library using a yeast two-hybrid system.". Hypertens. Res. 25 (4): 647–52. doi:10.1291/hypres.25.647. PMID 12358155.
- Strausberg RL, Feingold EA, Grouse LH, et al. (2003). "Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences.". Proc. Natl. Acad. Sci. U.S.A. 99 (26): 16899–903. doi:10.1073/pnas.242603899. PMC 139241
 . PMID 12477932.
. PMID 12477932. - Deloukas P, Earthrowl ME, Grafham DV, et al. (2004). "The DNA sequence and comparative analysis of human chromosome 10.". Nature. 429 (6990): 375–81. doi:10.1038/nature02462. PMID 15164054.
- Gerhard DS, Wagner L, Feingold EA, et al. (2004). "The status, quality, and expansion of the NIH full-length cDNA project: the Mammalian Gene Collection (MGC).". Genome Res. 14 (10B): 2121–7. doi:10.1101/gr.2596504. PMC 528928
 . PMID 15489334.
. PMID 15489334. - Bruneel A, Labas V, Mailloux A, et al. (2006). "Proteomics of human umbilical vein endothelial cells applied to etoposide-induced apoptosis.". Proteomics. 5 (15): 3876–84. doi:10.1002/pmic.200401239. PMID 16130169.
- Ewing RM, Chu P, Elisma F, et al. (2007). "Large-scale mapping of human protein-protein interactions by mass spectrometry.". Mol. Syst. Biol. 3 (1): 89. doi:10.1038/msb4100134. PMC 1847948
 . PMID 17353931.
. PMID 17353931. - Takahashi M, Watari E, Shinya E, et al. (2007). "Suppression of virus replication via down-modulation of mitochondrial short chain enoyl-CoA hydratase in human glioblastoma cells.". Antiviral Res. 75 (2): 152–8. doi:10.1016/j.antiviral.2007.02.002. PMID 17395278.