Terminator (genetics)
In genetics, a transcription terminator is a section of nucleic acid sequence that marks the end of a gene or operon in genomic DNA during transcription. This sequence mediates transcriptional termination by providing signals in the newly synthesized mRNA that trigger processes which release the mRNA from the transcriptional complex. These processes include the direct interaction of the mRNA secondary structure with the complex and/or the indirect activities of recruited termination factors. Release of the transcriptional complex frees RNA polymerase and related transcriptional machinery to begin transcription of new mRNAs.
Terminators in prokaryotes
Two classes of transcription terminators, Rho-dependent and Rho-independent, have been identified throughout prokaryotic genomes. These widely distributed sequences are responsible for triggering the end of transcription upon normal completion of gene or operon transcription, mediating early termination of transcripts as a means of regulation such as that observed in transcriptional attenuation, and to ensure the termination of runaway transcriptional complexes that manage to escape earlier terminators by chance, which prevents unnecessary energy expenditure for the cell.
Rho-dependent terminators
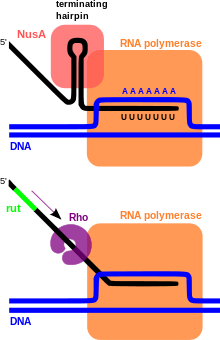
Rho-dependent transcription terminators require a protein called Rho factor, which exhibits RNA helicase activity, to disrupt the mRNA-DNA-RNA polymerase transcriptional complex. Rho-dependent terminators are found in bacteria and phage. The Rho-dependent terminator occurs downstream of translational stop codons and consists of an unstructured, cytosine-rich sequence on the mRNA known as a Rho utilization site (rut) for which a consensus sequence has not been identified, and a downstream transcription stop point (tsp). The rut serves as a mRNA loading site and as an activator for Rho; activation enables Rho to efficiently hydrolyze ATP and translocate down the mRNA while it maintains contact with the rut site. Rho is able to catch up with the RNA polymerase, which is stalled at the downstream tsp sites.[1] Contact between Rho and the RNA polymerase complex stimulates dissociation of the transcriptional complex through a mechanism involving allosteric effects of Rho on RNA polymerase.[2][3]
Rho-independent terminators
Intrinsic transcription terminators or Rho-independent terminators require the formation of a self-annealing hairpin structure on the elongating transcript, which results in the disruption of the mRNA-DNA-RNA polymerase ternary complex. The terminator sequence in DNA contains a 20 basepair GC-rich region of dyad symmetry followed by a short poly-T tract or "T stretch" which is transcribed to form the terminating hairpin and a 7-9 nucleotide "U tract" respectively. The mechanism of termination is hypothesized to occur through a combination of direct promotion of dissociation through allosteric effects of hairpin binding interactions with the RNA polymerase and "competitive kinetics". The hairpin formation causes RNA polymerase stalling and destabilization, leading to a greater likelihood that dissociation of the complex will occur at that location due to an increased time spent paused at that site and reduced stability of the complex.[4][5] Additionally, the elongation protein factor NusA interacts with the RNA polymerase and the hairpin structure to stimulate transcriptional termination.[6]
Terminators in eukaryotes
In eukaryotic transcription of mRNAs, terminator signals are recognized by protein factors that are associated with the RNA polymerase II and which trigger the termination process. Once the poly-A signals are transcribed into the mRNA, the proteins cleavage and polyadenylation specificity factor (CPSF) and cleavage stimulation factor (CstF) transfer from the carboxyl terminal domain of RNA polymerase II to the poly-A signal. These two factors then recruit other proteins to the site to cleave the transcript, freeing the mRNA from the transcription complex, and add a string of about 200 A-repeats to the 3' end of the mRNA in a process known as polyadenylation. During these processing steps, the RNA polymerase continues to transcribe for several kilobases and eventually dissociates from the DNA and downstream transcript through an unclear mechanism; there are two basic models for this event known as the torpedo and allosteric models.[7]
Torpedo model
After the mRNA is completed, the residual RNA strand remains in association with the DNA template and the RNA polymerase II, continuing to be transcribed. XRN2 (5'-3' Exoribonuclease 2), a RNase, attaches to the carboxyl terminal domain of RNA polymerase II and proceeds to degrade the uncapped residual RNA from 5’ to 3’ until it reaches the RNA pol II. 5' cap refers to a modified guanine added to the front of mRNA for protection from RNase. 3' poly(A) tail is added to the end of a mRNA strand for protection from exonucleases. Similar to Rho-dependent termination, XRN2 triggers dissociation of RNA polymerase II by either pushing the polymerase off of the DNA template or pulling the template out of the RNA polymerase.[8] The entire mechanism remains unclear.
Allosteric model
RNA polymerase normally is capable of transcribing DNA into single-stranded mRNA efficiently. However, upon transcribing over the poly-A signals on the DNA template, a conformational shift is induced in the RNA polymerase from the proposed loss of associated proteins from its carboxyl terminal domain. This change of conformation reduces RNA polymerase's processivity making the enzyme more prone to dissociating from its DNA-RNA substrate. In this case, termination is not completed by degradation of mRNA but instead is mediated by limiting the elongation efficiency of RNA polymerase and thus increasing the likelihood that the polymerase will dissociate and end its current cycle of transcription.[7]
See also
References
- ↑ Richardson, J. P. (1996). "Rho-dependent Termination of Transcription Is Governed Primarily by the Upstream Rho Utilization (rut) Sequences of a Terminator". Journal of Biological Chemistry. 271 (35): 21597–21603. doi:10.1074/jbc.271.35.21597. ISSN 0021-9258.
- ↑ Ciampi, MS. (Sep 2006). "Rho-dependent terminators and transcription termination". Microbiology. 152 (Pt 9): 2515–28. doi:10.1099/mic.0.28982-0. PMID 16946247.
- ↑ Epshtein, V; Dutta, D; Wade, J; Nudler, E (Jan 14, 2010). "An allosteric mechanism of Rho-dependent transcription termination.". Nature. 463 (7278): 245–9. doi:10.1038/nature08669. PMC 2929367
 . PMID 20075920.
. PMID 20075920. - ↑ von Hippel, P. H. (1998). "An Integrated Model of the Transcription Complex in Elongation, Termination, and Editing". Science. 281 (5377): 660–665. doi:10.1126/science.281.5377.660.
- ↑ Gusarov, Ivan; Nudler, Evgeny (1999). "The Mechanism of Intrinsic Transcription Termination". Molecular Cell. 3 (4): 495–504. doi:10.1016/S1097-2765(00)80477-3. ISSN 1097-2765.
- ↑ Santangelo, TJ.; Artsimovitch, I. (May 2011). "Termination and antitermination: RNA polymerase runs a stop sign.". Nat Rev Microbiol. 9 (5): 319–29. doi:10.1038/nrmicro2560. PMC 3125153
 . PMID 21478900.
. PMID 21478900. - 1 2 Watson, J. (2008). Molecular Biology of the Gene. Cold Spring Harbor Laboratory Press. pp. 410–411. ISBN 978-0-8053-9592-1.
- ↑ Luo, W.; Bartley D. (2004). "A ribonucleolytic rat torpedoes RNA polymerase II". Cell. 119 (7): 911–914. doi:10.1016/j.cell.2004.11.041. PMID 15620350.
External links
- Terminator Sequence at the US National Library of Medicine Medical Subject Headings (MeSH)