Glycoside hydrolase family 27
| Melibiase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
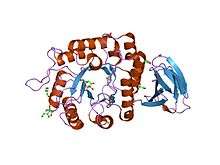 crystal structure of rice alpha-galactosidase | |||||||||
| Identifiers | |||||||||
| Symbol | Melibiase | ||||||||
| Pfam | PF02065 | ||||||||
| Pfam clan | CL0058 | ||||||||
| InterPro | IPR000111 | ||||||||
| SCOP | 1ktc | ||||||||
| SUPERFAMILY | 1ktc | ||||||||
| CAZy | GH27 | ||||||||
| |||||||||
In molecular biology, glycoside hydrolase family 27 is a family of glycoside hydrolases.
Glycoside hydrolases EC 3.2.1. are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A classification system for glycoside hydrolases, based on sequence similarity, has led to the definition of >100 different families.[1][2][3] This classification is available on the CAZy (http://www.cazy.org) web site,[4] and also discussed at CAZypedia, an online encyclopedia of carbohydrate active enzymes.[5]
Glycoside hydrolase family 27 together with family 31 and the family 36 alpha-galactosidases form the glycosyl hydrolase clan GH-D (CAZY GH), a superfamily of alpha-galactosidases, alpha-N-acetylgalactosaminidases, and isomaltodextranases which are likely to share a common catalytic mechanism and structural topology.
Alpha-galactosidase (EC 3.2.1.22) (melibiase)[6] catalyzes the hydrolysis of melibiose into galactose and glucose. In man, the deficiency of this enzyme is the cause of Fabry's disease (X-linked sphingolipidosis). Alpha-galactosidase is present in a variety of organisms. There is a considerable degree of similarity in the sequence of alpha-galactosidase from various eukaryotic species. Escherichia coli alpha-galactosidase (gene melA), which requires NAD and magnesium as cofactors, is not structurally related to the eukaryotic enzymes; by contrast, an Escherichia coli plasmid encoded alpha-galactosidase (gene rafA P16551)[7] contains a region of about 50 amino acids which is similar to a domain of the eukaryotic alpha-galactosidases. Alpha-N-acetylgalactosaminidase (EC 3.2.1.49)[8] catalyzes the hydrolysis of terminal non-reducing N-acetyl-D-galactosamine residues in N-acetyl-alpha-D- galactosaminides. In man, the deficiency of this enzyme is the cause of Schindler and Kanzaki diseases. The sequence of this enzyme is highly related to that of the eukaryotic alpha-galactosidases.
References
- ↑ Henrissat B, Callebaut I, Mornon JP, Fabrega S, Lehn P, Davies G (1995). "Conserved catalytic machinery and the prediction of a common fold for several families of glycosyl hydrolases". Proc. Natl. Acad. Sci. U.S.A. 92 (15): 7090–7094. doi:10.1073/pnas.92.15.7090. PMC 41477
 . PMID 7624375.
. PMID 7624375. - ↑ Henrissat B, Davies G (1995). "Structures and mechanisms of glycosyl hydrolases". Structure. 3 (9): 853–859. doi:10.1016/S0969-2126(01)00220-9. PMID 8535779.
- ↑ Bairoch, A. "Classification of glycosyl hydrolase families and index of glycosyl hydrolase entries in SWISS-PROT". 1999.
- ↑ Henrissat, B. and Coutinho P.M. "Carbohydrate-Active Enzymes server". 1999.
- ↑ CAZypedia, an online encyclopedia of carbohydrate-active enzymes.
- ↑ Dey PM, Pridham JB (1972). "Biochemistry of alpha-galactosidases". Adv. Enzymol. Relat. Areas Mol. Biol. 36: 91–120. PMID 4561015.
- ↑ Aslanidis C, Schmid K, Schmitt R (1989). "Nucleotide sequences and operon structure of plasmid-borne genes mediating uptake and utilization of raffinose in Escherichia coli". J. Bacteriol. 171 (12): 6753–6763. PMC 210573
 . PMID 2556373.
. PMID 2556373. - ↑ Wang AM, Bishop DF, Desnick RJ (1990). "Human alpha-N-acetylgalactosaminidase-molecular cloning, nucleotide sequence, and expression of a full-length cDNA. Homology with human alpha-galactosidase A suggests evolution from a common ancestral gene". J. Biol. Chem. 265 (35): 21859–21866. PMID 2174888.
This article incorporates text from the public domain Pfam and InterPro IPR000111