Western blot

The western blot (sometimes called the protein immunoblot) is a widely used analytical technique used in molecular biology, immunogenetics and other molecular biology disciplines to detect specific proteins in a sample of tissue homogenate or extract.
Artificial antibodies are created that react with a specific target protein. The sample to be tested is prepared and put together with these antibodies on a membrane – if the specific protein sought for is present, after a gel electrophoresis step this will result in an accordingly stained band on the western blot.
A number of search engines, such as CiteAb, Antibodypedia, and SeekProducts, are available that can help researchers find suitable antibodies for use in western blotting. Many reagent companies specialize in providing antibodies (both monoclonal and polyclonal antibodies) against tens of thousands of different proteins.[1]
Other related techniques include dot blot analysis, immunohistochemistry and immunocytochemistry where antibodies are used to detect proteins in tissues and cells by immunostaining, and enzyme-linked immunosorbent assay (ELISA).
The name western blot is a play on the eponymously-named Southern blot, a technique for DNA detection developed earlier by Edwin Southern. Similarly, detection of RNA is termed northern blot and was developed by George Stark at Stanford.[2] The term "western blot" was given to the technique by W. Neal Burnette,[3] although the method itself originated in the laboratory of Harry Towbin at the Friedrich Miescher Institute.[4]
Applications
- The confirmatory HIV test employs a western blot to detect anti-HIV antibody in a human serum sample. Proteins from known HIV-infected cells are separated and blotted on a membrane as above. Then, the serum to be tested is applied in the primary antibody incubation step; free antibody is washed away, and a secondary anti-human antibody linked to an enzyme signal is added. The stained bands then indicate the proteins to which the patient's serum contains antibody.
- A western blot is also used as the definitive test for bovine spongiform encephalopathy (BSE, commonly referred to as 'mad cow disease').
- Some forms of Lyme disease testing employ western blotting.
- A western blot can also be used as a confirmatory test for Hepatitis B infection and HSV-2 (Herpes Type 2) infection.
- In veterinary medicine, a western blot is sometimes used to confirm FIV+ status in cats.
Procedure
To create a western blot a gel electrophoresis is used to separate native proteins by 3-D structure or denatured proteins by the length of the polypeptide. The proteins are then transferred to a membrane (typically nitrocellulose or PVDF), where they are stained with antibodies specific to the target protein.[4][5] The gel electrophoresis step is included in western blot analysis to resolve the issue of the cross-reactivity of antibodies.
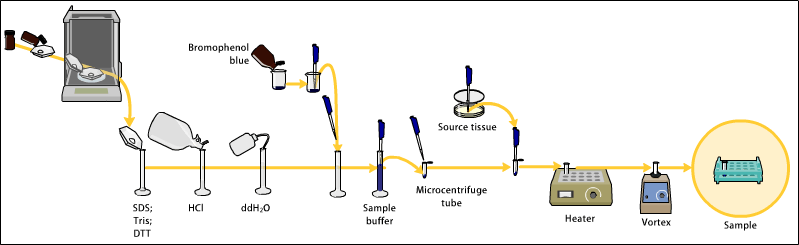
Tissue preparation
Samples can be taken from whole tissue or from cell culture. Solid tissues are first broken down mechanically using a blender (for larger sample volumes), using a homogenizer (smaller volumes), or by sonication. Cells may also be broken open by one of the above mechanical methods. However, virus or environmental samples can be the source of protein and thus western blotting is not restricted to cellular studies only.
Assorted detergents, salts, and buffers may be employed to encourage lysis of cells and to solubilize proteins. Protease and phosphatase inhibitors are often added to prevent the digestion of the sample by its own enzymes. Tissue preparation is often done at cold temperatures to avoid protein denaturing and degradation.
A combination of biochemical and mechanical techniques – comprising various types of filtration and centrifugation – can be used to separate different cell compartments and organelles.
Gel electrophoresis
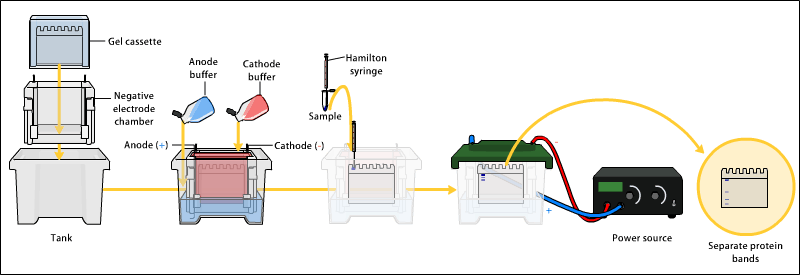
The proteins of the sample are separated using gel electrophoresis. Separation of proteins may be by isoelectric point (pI), molecular weight, electric charge, or a combination of these factors. The nature of the separation depends on the treatment of the sample and the nature of the gel. This is a very useful way to identify a protein.
By far the most common type of gel electrophoresis employs polyacrylamide gels and buffers loaded with sodium dodecyl sulfate (SDS). SDS-PAGE (SDS polyacrylamide gel electrophoresis) maintains polypeptides in a denatured state once they have been treated with strong reducing agents to remove secondary and tertiary structure (e.g. disulfide bonds [S-S] to sulfhydryl groups [SH and SH]) and thus allows separation of proteins by their molecular weight. Sampled proteins become covered in the negatively charged SDS and move to the positively charged electrode through the acrylamide mesh of the gel. Smaller proteins migrate faster through this mesh, and the proteins are thus separated according to size (usually measured in kilodaltons, kDa). The concentration of acrylamide determines the resolution of the gel - the greater the acrylamide concentration, the better the resolution of lower molecular weight proteins. The lower the acrylamide concentration, the better the resolution of higher molecular weight proteins. Proteins travel only in one dimension along the gel for most blots.
Samples are loaded into wells in the gel. One lane is usually reserved for a marker or ladder, a commercially available mixture of proteins having defined molecular weights, typically stained so as to form visible, coloured bands. When voltage is applied along the gel, proteins migrate through it at different speeds dependent on their size. These different rates of advancement (different electrophoretic mobilities) separate into bands within each lane.
It is also possible to use a two-dimensional (2-D) gel which spreads the proteins from a single sample out in two dimensions. Proteins are separated according to isoelectric point (pH at which they have a neutral net charge) in the first dimension, and according to their molecular weight in the second dimension.
Transfer
To make the proteins accessible to antibody detection, they are moved from within the gel onto a membrane made of nitrocellulose or polyvinylidene difluoride (PVDF). The primary method for transferring the proteins is called electroblotting and uses an electric current to pull proteins from the gel into the PVDF or nitrocellulose membrane. The proteins move from within the gel onto the membrane while maintaining the organization they had within the gel. An older method of transfer involves placing a membrane on top of the gel, and a stack of filter papers on top of that. The entire stack is placed in a buffer solution which moves up the paper by capillary action, bringing the proteins with it. In practice this method is not used as it takes too much time; electroblotting is preferred. As a result of either "blotting" process, the proteins are exposed on a thin surface layer for detection (see below). Both varieties of membrane are chosen for their non-specific protein binding properties (i.e. binds all proteins equally well). Protein binding is based upon hydrophobic interactions, as well as charged interactions between the membrane and protein. Nitrocellulose membranes are cheaper than PVDF, but are far more fragile and do not stand up well to repeated probings.
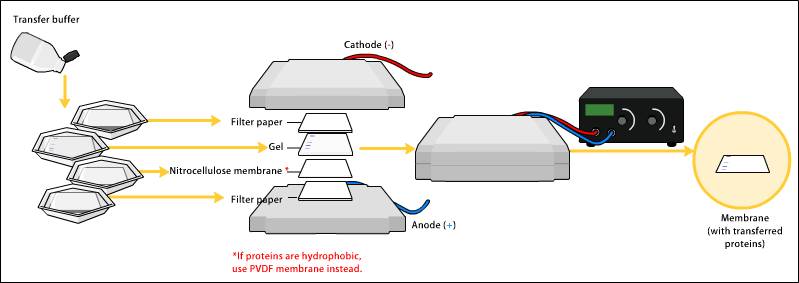
The uniformity and overall effectiveness of transfer of protein from the gel to the membrane can be checked by staining the membrane with Coomassie Brilliant Blue or Ponceau S dyes. Ponceau S is the more common of the two, due to its higher sensitivity and water solubility, the latter making it easier to subsequently destain and probe the membrane, as described below.[6]
Blocking
Since the membrane has been chosen for its ability to bind protein and as both antibodies and the target are proteins, steps must be taken to prevent the interactions between the membrane and the antibody used for detection of the target protein. Blocking of non-specific binding is achieved by placing the membrane in a dilute solution of protein - typically 3-5% Bovine serum albumin (BSA) or non-fat dry milk (both are inexpensive) in Tris-Buffered Saline (TBS) or I-Block, with a minute percentage (0.1%) of detergent such as Tween 20 or Triton X-100. The protein in the dilute solution attaches to the membrane in all places where the target proteins have not attached. Thus, when the antibody is added, there is no room on the membrane for it to attach other than on the binding sites of the specific target protein. This reduces background in the final product of the western blot, leading to clearer results, and eliminates false positives.
Incubation
During the detection process the membrane is "probed" for the protein of interest with a modified antibody which is linked to a reporter enzyme; when exposed to an appropriate substrate, this enzyme drives a colorimetric reaction and produces a color. For a variety of reasons, this traditionally takes place in a two-step process, although there are now one-step detection methods available for certain applications.
Two steps
The primary antibodies are generated when a host species or immune cell culture is exposed to the protein of interest (or a part thereof). Normally, this is part of the immune response, whereas here they are harvested and used as sensitive and specific detection tools that bind the protein directly.
After blocking, a dilute solution of primary antibody (generally between 0.5 and 5 micrograms/mL) is incubated with the membrane under gentle agitation. Typically, the solution is composed of buffered saline solution with a small percentage of detergent, and sometimes with powdered milk or BSA. The antibody solution and the membrane can be sealed and incubated together for anywhere from 30 minutes to overnight. It can also be incubated at different temperatures, with lesser temperatures being associated with more binding, both specific (to the target protein, the "signal") and non-specific ("noise").
After rinsing the membrane to remove unbound primary antibody, the membrane is exposed to another antibody, directed at a species-specific portion of the primary antibody. Antibodies come from animal sources (or animal sourced hybridoma cultures); an anti-mouse secondary will bind to almost any mouse-sourced primary antibody, which allows some cost savings by allowing an entire lab to share a single source of mass-produced antibody, and provides far more consistent results. This is known as a secondary antibody, and due to its targeting properties, tends to be referred to as "anti-mouse," "anti-goat," etc. The secondary antibody is usually linked to biotin or a reporter enzyme such as alkaline phosphatase or horseradish peroxidase. This means that several secondary antibodies will bind to one primary antibody and enhance the signal.
Most commonly, a horseradish peroxidase-linked secondary is used to cleave a chemiluminescent agent, and the reaction product produces luminescence in proportion to the amount of protein. A sensitive sheet of photographic film is placed against the membrane, and exposure to the light from the reaction creates an image of the antibodies bound to the blot. A cheaper but less sensitive approach utilizes a 4-chloronaphthol stain with 1% hydrogen peroxide; the reaction of peroxide radicals with 4-chloronaphthol produces a dark purple stain that can be photographed without using specialized photographic film.
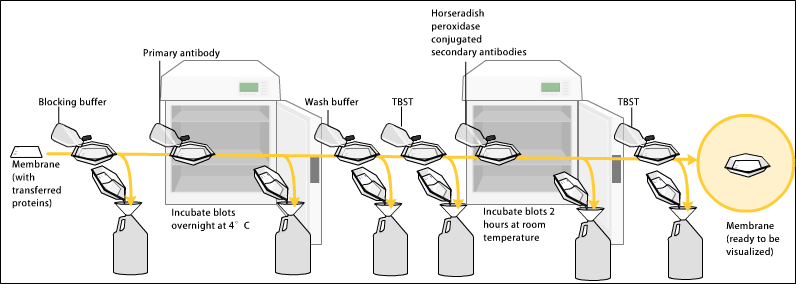
As with the ELISPOT and ELISA procedures, the enzyme can be provided with a substrate molecule that will be converted by the enzyme to a colored reaction product that will be visible on the membrane (see the figure below with blue bands).
Another method of secondary antibody detection utilizes a near-infrared (NIR) fluorophore-linked antibody. The light produced from the excitation of a fluorescent dye is static, making fluorescent detection a more precise and accurate measure of the difference in the signal produced by labeled antibodies bound to proteins on a western blot. Proteins can be accurately quantified because the signal generated by the different amounts of proteins on the membranes is measured in a static state, as compared to chemiluminescence, in which light is measured in a dynamic state.[7]
A third alternative is to use a radioactive label rather than an enzyme coupled to the secondary antibody, such as labeling an antibody-binding protein like Staphylococcus Protein A or Streptavidin with a radioactive isotope of iodine. Since other methods are safer, quicker, and cheaper, this method is now rarely used; however, an advantage of this approach is the sensitivity of auto-radiography-based imaging, which enables highly accurate protein quantification when combined with optical software (e.g. Optiquant).
One step
Historically, the probing process was performed in two steps because of the relative ease of producing primary and secondary antibodies in separate processes. This gives researchers and corporations huge advantages in terms of flexibility, and adds an amplification step to the detection process. Given the advent of high-throughput protein analysis and lower limits of detection, however, there has been interest in developing one-step probing systems that would allow the process to occur faster and with fewer consumables. This requires a probe antibody which both recognizes the protein of interest and contains a detectable label, probes which are often available for known protein tags. The primary probe is incubated with the membrane in a manner similar to that for the primary antibody in a two-step process, and then is ready for direct detection after a series of wash steps.
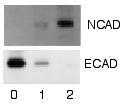
Detection / Visualization
After the unbound probes are washed away, the western blot is ready for detection of the probes that are labeled and bound to the protein of interest. In practical terms, not all westerns reveal protein only at one band in a membrane. Size approximations are taken by comparing the stained bands to that of the marker or ladder loaded during electrophoresis. The process is commonly repeated for a structural protein, such as actin or tubulin, that should not change between samples. The amount of target protein is normalized to the structural protein to control between groups. A superior strategy is the normalization to the total protein visualized with trichloroethanol[8][9] or epicocconone.[10] This practice ensures correction for the amount of total protein on the membrane in case of errors or incomplete transfers. (see western blot normalization)
Colorimetric detection
The colorimetric detection method depends on incubation of the western blot with a substrate that reacts with the reporter enzyme (such as peroxidase) that is bound to the secondary antibody. This converts the soluble dye into an insoluble form of a different color that precipitates next to the enzyme and thereby stains the membrane. Development of the blot is then stopped by washing away the soluble dye. Protein levels are evaluated through densitometry (how intense the stain is) or spectrophotometry.
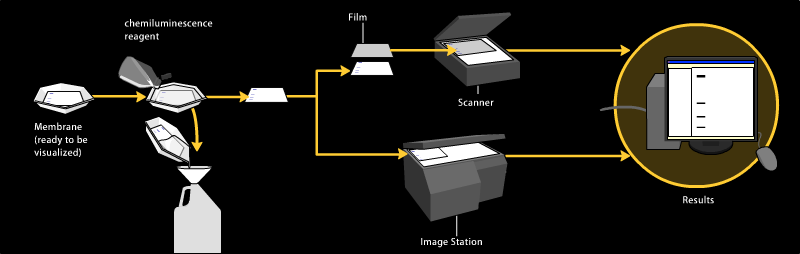
Chemiluminescent detection
Chemiluminescent detection methods depend on incubation of the western blot with a substrate that will luminesce when exposed to the reporter on the secondary antibody. The light is then detected by CCD cameras which capture a digital image of the western blot or photographic film. The use of film for western blot detection is slowly disappearing because of non linearity of the image (non accurate quantification). The image is analysed by densitometry, which evaluates the relative amount of protein staining and quantifies the results in terms of optical density. Newer software allows further data analysis such as molecular weight analysis if appropriate standards are used.
The main companies offering Chemiluminescence imaging systems are Analytik Jena, Syngene, UVP, Azure Biosystems, UVITEC, GE, Biorad and Vilber Lourmat.
Radioactive detection
Radioactive labels do not require enzyme substrates, but rather, allow the placement of medical X-ray film directly against the western blot, which develops as it is exposed to the label and creates dark regions which correspond to the protein bands of interest (see image to the right). The importance of radioactive detections methods is declining due to its hazardous radiation , because it is very expensive, health and safety risks are high, and ECL (enhanced chemiluminescence) provides a useful alternative.
Fluorescent detection
The fluorescently labeled probe is excited by light and the emission of the excitation is then detected by a photosensor such as a CCD camera equipped with appropriate emission filters which captures a digital image of the western blot and allows further data analysis such as molecular weight analysis and a quantitative western blot analysis. Fluorescence is considered to be one of the best methods for quantification but is less sensitive than chemiluminescence.[11]
Secondary probing
One major difference between nitrocellulose and PVDF membranes relates to the ability of each to support "stripping" antibodies off and reusing the membrane for subsequent antibody probes. While there are well-established protocols available for stripping nitrocellulose membranes, the sturdier PVDF allows for easier stripping, and for more reuse before background noise limits experiments. Another difference is that, unlike nitrocellulose, PVDF must be soaked in 95% ethanol, isopropanol or methanol before use. PVDF membranes also tend to be thicker and more resistant to damage during use.
2-D gel electrophoresis
2-dimensional SDS-PAGE uses the principles and techniques outlined above. 2-D SDS-PAGE, as the name suggests, involves the migration of polypeptides in 2 dimensions. For example, in the first dimension, polypeptides are separated according to isoelectric point, while in the second dimension, polypeptides are separated according to their molecular weight. The isoelectric point of a given protein is determined by the relative number of positively (e.g. lysine, arginine) and negatively (e.g. glutamate, aspartate) charged amino acids, with negatively charged amino acids contributing to a low isoelectric point and positively charged amino acids contributing to a high isoelectric point. Samples could also be separated first under nonreducing conditions using SDS-PAGE, and under reducing conditions in the second dimension, which breaks apart disulfide bonds that hold subunits together. SDS-PAGE might also be coupled with urea-PAGE for a 2-dimensional gel.
In principle, this method allows for the separation of all cellular proteins on a single large gel. A major advantage of this method is that it often distinguishes between different isoforms of a particular protein - e.g. a protein that has been phosphorylated (by addition of a negatively charged group). Proteins that have been separated can be cut out of the gel and then analysed by mass spectrometry, which identifies the protein.
See also
- Far-Eastern blotting
- Far-western blotting
- Eastern blot
- Northwestern blot
- SDS-PAGE
- Fast parallel proteolysis (FASTpp)
References
- ↑ "Western blot antibody". exactantigen.com. Retrieved 2009-01-29.
- ↑ Alwine J, Kemp D, Stark G (1977). "Method for detection of specific RNAs in agarose gels by transfer to diazobenzyloxymethyl-paper and hybridization with DNA probes.". Proceedings of the National Academy of Sciences USA. 74 (12): 5350–5354. doi:10.1073/pnas.74.12.5350. PMC 431715
 . PMID 414220.
. PMID 414220. - ↑ Burnette WN. (1981). "'Western blotting': electrophoretic transfer of proteins from sodium dodecyl sulfate—polyacrylamide gels to unmodified nitrocellulose and radiographic detection with antibody and radioiodinated protein A". Analytical Biochemistry. 112 (2): 195–203. doi:10.1016/0003-2697(81)90281-5. ISSN 0003-2697. PMID 6266278.
- 1 2 Towbin H, Staehelin T, Gordon J (1979). "Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications". Proceedings of the National Academy of Sciences USA. 76 (9): 4350–54. doi:10.1073/pnas.76.9.4350. PMC 411572
 . PMID 388439.
. PMID 388439. - ↑ Renart J, Reiser J, Stark GR (1979). "Transfer of proteins from gels to diazobenzyloxymethyl-paper and detection with antisera: a method for studying antibody specificity and antigen structure". Proceedings of the National Academy of Sciences USA. 76 (7): 3116–20. doi:10.1073/pnas.76.7.3116. PMC 383774
 . PMID 91164.
. PMID 91164. - ↑ Corley RB. (2005). A Guide to Methods in the Biomedical Sciences. Springer. p. 11. ISBN 978-0-387-22844-0.
- ↑ Ambroz K. (2006-09-20). "Improving quantification accuracy for western blots" (PDF). Image Analysis.
- ↑ Stennert, E; Arold, R (1973). "The double external auditory canal (author's transl)". HNO. 21 (10): 293–6. PMID 4769330.
- ↑ Gilda, J. E.; Gomes, A. V. (2013). "Stain-Free total protein staining is a superior loading control to β-actin for Western blots". Analytical Biochemistry. 440 (2): 186–8. doi:10.1016/j.ab.2013.05.027. PMC 3809032
 . PMID 23747530.
. PMID 23747530. - ↑ Moritz, C. P.; Marz, S. X.; Reiss, R; Schulenborg, T; Friauf, E (2014). "Epicocconone staining: A powerful loading control for Western blots". Proteomics. 14 (2–3): 162–8. doi:10.1002/pmic.201300089. PMID 24339236.
- ↑ "Imaging Systems for Westerns: Chemiluminescence vs. Infrared Detection, 2009 - , Methods in Molecular Biology, Protein Blotting and Detection, vol. 536" (PDF). Humana Press. Retrieved 2010-02-26.
External links
| Library resources about Western immunoblotting |
- Western blot analysis of sub-cellular fractionated samples using the Odyssey Infrared Imaging System (a protocol)
- Western blot protocol including example buffers and reprobing procedure
- Western blot protocol including troubleshooting guide
- Image Studio Lite Western Blot Analysis Software (Free)
- Western Blotting Principles and Methods Handbook
- Advansta WiKi on Western Blots
Protocols
- Western Blot Video Protocol
- Western Blotting Protocol from Protocolmonkey
- Modified western blotting protocols from Biotechniques
- Dr. Mark Barton Frank Lab protocol
- [ Kamps's Western Blotting Protocol]
- CSH Protocols collection: Immunoblotting
- Western Blot Protocol from HowtoWesternBlot.net
- https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3456489/