Spindle checkpoint
During the process of cell division, the spindle checkpoint prevents separation of the duplicated chromosomes until each chromosome is properly attached to the spindle apparatus. In order to preserve the cell's identity and proper function, it is necessary to maintain the appropriate number of chromosomes after each cell division. An error in generating daughter cells with fewer or greater number of chromosomes than expected (a situation termed aneuploidy), may lead in best case to cell death, or alternatively it may generate catastrophic phenotypic results. Examples include:
- In cancer cells, aneuploidy is a frequent event, indicating that these cells present a defect in the machinery involved in chromosome segregation, as well as in the mechanism ensuring that segregation is correctly performed.
- In humans, Down syndrome appears in children carrying in their cells one extra copy of chromosome 21, as a result of a defect in chromosome segregation during meiosis in one of the progenitors. This defect will generate a gamete (spermatozoide or oocyte) with an extra chromosome 21. After fecundation, this gamete will generate an embryo with three copies of chromosome 21.
The mechanisms verifying that all the requirements to pass to the next phase in the cell cycle have been fulfilled are called checkpoints. All along the cell cycle, there are different checkpoints. The checkpoint ensuring that chromosome segregation is correct is termed spindle assembly checkpoint (SAC), spindle checkpoint or mitotic checkpoint. During mitosis or meiosis, the spindle checkpoint prevents anaphase onset until all chromosomes are properly attached to the spindle. To achieve proper segregation, the two kinetochores on the sister chromatids must be attached to opposite spindle poles (bipolar orientation). Only this pattern of attachment will ensure that each daughter cell receives one copy of the chromosome.
Chromosome segregation
Cell division: duplication of material and distribution to daughter cells
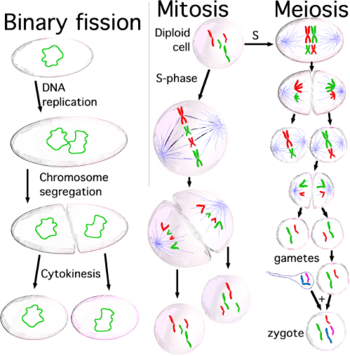
When cells are ready to divide, because cell size is big enough or because they receive the appropriate stimulus,[1] they activate the mechanism to enter into the cell cycle, and they duplicate most organelles during S (synthesis) phase, including their centrosome. Therefore, when the cell division process will end, each daughter cell will receive a complete set of organelles. At the same time, during S phase all cells must duplicate their DNA very precisely, a process termed DNA replication. Once DNA replication has finished, in eukaryotes the DNA molecule is compacted and condensed, to form the mitotic chromosomes, each one constituted by two sister chromatids, which stay held together by the establishment of cohesion between them; each chromatid is a complete DNA molecule, attached via microtubules to one of the two centrosomes of the dividing cell, located at opposed poles of the cell. The structure formed by the centrosomes and the microtubules is named mitotic spindle, due to its characteristic shape, holding the chromosomes between the two centrosomes. Both sister chromatids stay together until anaphase; at this moment they separate from each other and they travel towards the centrosome to which they are attached. In this way, when the two daughter cells separate at the end of the division process, each one will receive a complete set of chromatids. The mechanism responsible for the correct distribution of sister chromatids during cell division is named chromosome segregation.
To ensure that chromosome segregation takes place correctly, cells have developed a precise and complex mechanism. In the first place, cells must coordinate centrosome duplication with DNA replication, and a failure in this coordination will generate monopolar or multipolar mitotic spindles, which generally will produce abnormal chromosome segregation,[2] because in this case, chromosome distribution will not take place in a balanced way.
Mitosis: anchoring of chromosomes to the spindle and chromosome segregation
During S phase, the centrosome starts to duplicate. Just at the beginning of mitosis, both centrioles achieve their maximal length, recruit additional material and their capacity to nucleate microtubules increases. As mitosis progresses, both centrosomes separate to generate the mitotic spindle.[3] In this way, the mitotic spindle has two poles emanating microtubules. Microtubules (MTs) are long proteic filaments, with asymmetric extremities: one end termed "minus" (-) end, relatively stable and close to the centrosome, and an end termed "plus" (+) end, with alterning phases of growing-retraction, exploring the center of the cell searching the chromosomes. Each chromatid has a special region, named the centromere, on top of which is assembled a proteic structure termed kinetochore, which is able to stabilize the microtubule plus end. Therefore, if by chance a microtubule exploring the center of the cell encounters a kinetochore, it may happen that the kinetochore will capture it, so that the chromosome will become attached to the spindle via the kinetochore of one of its sister chromatids. As it happens that sister chromatids are attached together and both kinetochores are located back-to-back on both chromatids, when one kinetochore becomes attached to one centrosome, the sister kinetochore becomes exposed to the centrosome located in the opposed pole; for this reason, in most cases the second kinetochore becomes associated to the centrosome in the opposed pole, via its microtubules,[4] so that the chromosomes become "bi-oriented", a fundamental configuration (also named amphitelic) to ensure that chromosome segregation will take place correctly when the cell will divide.[5][6] Occasionally, one of the two sister kinetochores may attach simultaneously to MTs generated by both poles, a configuration named merotelic, which is not detected by the spindle checkpoint but that may generate lagging chromosomes during anaphase and, consequently, aneuploidy. Merotelic orientation (characterized by the absence of tension between sister kinetochores) is frequent at the beginning of mitosis, but the protein Aurora B (a kinase conserved from yeast to vertebrates) detects and eliminates this type of anchoring.[7] (Note: Aurora B is frequently overexpressed in various types of tumors and currently is a target for the development of anticancer drugs.[8])
Discovery of the spindle assembly checkpoint (SAC)
Zirkle (in 1970) was one of the first researchers to observe that, when just one chromosome is retarded to arrive at the metaphase plate, anaphase onset is postponed until some minutes after its arrival.[9] This observation, together with similar ones, suggested that a control mechanism exists at the metaphase-to-anaphase transition. Using drugs such as nocodazole and colchicine, the mitotic spindle disassembles and the cell cycle is blocked at the metaphase-to-anaphase transition. Using these drugs (see the review from Rieder and Palazzo in 1992[10]), the putative control mechanism was named Spindle Assembly Checkpoint (SAC). This regulatory mechanism has been intensively studied since(see the review from Burke and Stukenberg in 2008[11]).
Using different types of genetic studies, it has been established that diverse kinds of defects are able to activate the SAC: spindle depolimerization,[12][13] the presence of dicentric chromosomes (with two centromeres),[14] centromeres segregating in an aberrant way,[15] defects in the spindle pole bodies in S. cerevisiae,[16] defects in the kinetochore proteins,[17] mutations in the centromeric DNA[18] or defects in the molecular motors active during mitosis.[12] A summary of these observations can be found in the article from Hardwick and collaborators in 1999.[19]
Using its own observations, Zirkle[9] was the first to propose that "some (…) susbstance, necessary for the cell to proceed to anaphase, appears some minutes after C (moment of the arrival of the last chromosome to the metaphase plate), or after a drastic change in the cytoplasmic condition, just at C or immediately after C", suggesting that this function is located on kinetochores unattached to the mitotic spindle. McIntosh extended this proposal, suggesting that one enzyme sensitive to tension located at the centromeres produces an inhibitor to the anaphase onset when the two sister kinetochores are not under bipolar tension.[20] Indeed, the available data suggested that the signal "wait to enter in anaphase" is produced mostly on or close to unattached kinetochores.[21] However, the primary event associated to the kinetochore attachment to the spindle, which is able to inactivate the inhibitory signal and release the metaphase arrest, could be either the acquisition of microtubules by the kinetochore (as proposed by Rieder and collaborators in 1995[21]), or the tension stabilizing the anchoring of microtubules to the kinetochores (as suggested by the experiments realized at Nicklas' lab[22]). Subsequent studies in cells containing two independent mitotic spindles in a sole cytoplasm showed that the inhibitor of the metaphase-to-anaphase transition is generated by unattached kinetochores and is not freely diffusible in the cytoplasm.[23] Yet in the same study it was shown that, once the transition from metaphase to anaphase is initiated in one part of the cell, this information is extended all along the cytoplasm, and can overcome the signal "wait to enter in anaphase" associated to a second spindle containing unattached kinetochores.
Sister chromatids cohesion during mitosis
Cohesin: SMC proteins
As it has been previously noted, sister chromatids stay associated from S phase (when DNA is replicated to generate two identic copies, the two chromatids) until anaphase. At this point, the two sister chromatids separate and travel to opposite poles in the dividing cell. Genetic and biochemical studies in yeast and in egg's extracts in Xenopus laevis identified a polyprotein complex as an essential player in sister chromatids cohesion (see the review from Hirano in 2000[24]). This complex is known as the cohesin complex and in Saccharomyces cerevisiae is composed of at least four subunits: Smc1p, Smc3p, Scc1p (or Mcd1p) and Scc3p. Both Smc1p and Smc3p belong to the family of proteins for the Structural Maintenance of Chromosomes (SMC), which constitute a group of chromosomic ATPases highly conserved, and form an heterodimer (Smc1p/Smc3p). Scc1p is the homolog in S.cerevisiae of Rad21, first identified as a protein involved in DNA repair in S. pombe. These four proteins are essential in yeast, and a mutation in any of them will produce premature sister chromatid separation. In yeast, cohesin binds to preferential sites along chromosome arms, and is very abundant close to the centromeres, as it was shown in a study using chromatin immunoprecipitation.[25]
The role of heterochromatin
Classical cytologic observations suggested that sister chromatids are more strongly attached at heterochromatic regions,[26] and this suggested that the special structure or composition of heterochromatin might favour cohesin recruitment.[27] In fact, it has been shown that Swi6 (the homolog of HP-1 in S. pombe) binds to methylated Lys 9 of histone H3 and promotes the binding of cohesin to the centromeric repeats in S. pombe.[28][29] More recent studies indicate that the RNAi machinery regulates heterochromatin establishment, which in turn recruits cohesin to this region, both in S. pombe[30] and in vertebrate cells.[31] However, there must be other mechanisms than heterochromatin to ensure an augmented cohesion at centromeres, because S. cerevisiae lacks heterochromatin next to centromeres, but the presence of a functional centromere induces an increase of cohesin association in a contiguous region, spanning 20-50kb.[32]
In this direction, Orc2 (one protein included in the origin recognition complex, ORC, implicated in the initiation of DNA replication during S phase) is also located on kinetochores during mitosis in human cells;[33] in agreement with this localization, some observations indicate that Orc2 in yeast is implicated in sister chromatid cohesion, and its removal induces SAC activation.[34] It has also been observed that other components of the ORC complex (such as orc5 in S. pombe) are implicated in cohesion.[35] However, the molecular pathway involving the ORC proteins seems to be additive to the cohesins' pathway, and it is mostly unknown.
Function of cohesion and its dissolution

Centromeric cohesion resists the forces exerted by spindle microtubules towards the poles, which generate tension between sister kinetochores. In turn, this tension stabilizes the attachment microtubule-kinetochore, through a mechanism implicating the protein Aurora B (a review about this issue : Hauf and Watanabe 2004[36]).
Indeed, a decrease in the cellular levels of cohesin generates the premature separation of sister chromatids, as well as defects in chromosome congression at the metaphase plate and delocalization of the proteins in the chromosomal passenger complex, which contains the protein Aurora B.[37][38] The proposed structure for the cohesin complex suggests that this complex connects directly both sister chromatids.[39] In this proposed structure, the SMC components of cohesin play a structural role, so that the SMC heterodimer may function as a DNA binding protein, whose conformation is regulated by ATP.[40] Scc1p and Scc3p, however, would play a regulatory role.[24]
In S. cerevisiae, Pds1p (also known as securin) regulates sister chromatids cohesion, because it binds and inhibits the protease Esp1p (separin or separase). When anaphase onset is triggered, the anaphase-promoting complex (APC/C or Cyclosome) degrades securin. Securin degradation releases the protease Esp1p/separase, which degrades the cohesin rings that link the two sister chromatids, therefore promoting sister chromatids separation.[41] It has been also shown that Polo/Cdc5 kinase phosphorylates serine residues next to the cutting site for Scc1, and this phosphorylation would facilitate the cutting activity.[42]
Although this machinery is conserved through evolution,[43][44] in vertebrates most cohesin molecules are released in prophase, independently of the presence of the APC/C, in a process dependent on Polo-like 1 (PLK1) and Aurora B.[45] Yet it has been shown that a small quantity of Scc1 remains associated to centromeres in human cells until metaphase, and a similar amount is cut in anaphase, when it disappears from centromeres.[46] On the other hand, some experiments show that sister chromatids cohesion in the arms is lost gradually after sister centromeres have separated, and sister chromatids move toward the opposite poles of the cell.[47][48]
According to some observations, a fraction of cohesins in the chromosomal arms and the centromeric cohesins are protected by the protein Shugoshin (Sgo1), avoiding their release during prophase.[49][50] To be able to function as protector for the centromeric cohesion, Sgo1 must be inactivated at the beginning of anaphase, as well as Pds1p. In fact, both Pds1p and Sgo1 are substrates of APC/C in vertebrates.[51]
Metaphase to anaphase transition
The beginning of metaphase is characterized by the connection of the microtubules to the kinetochores of the chromosomes, as well as the alignment of the chromosomes in the middle of the cell. Each chromatid has its own kinetochore, and all of the microtubules that are bound to kinetochores of sister chromatids radiate from opposite poles of the cell. These microtubules exert a pulling force on the chromosomes towards the opposite ends of the cells, while the cohesion between the sister chromatids opposes this force.
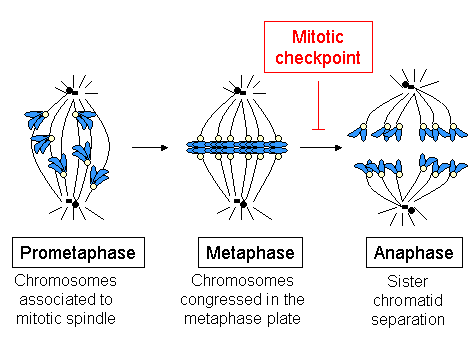
At the metaphase to anaphase transition, this cohesion between sister chromatids is dissolved, and the separated chromatids are pulled to opposite sides of the cell by the spindle microtubules. The chromatids are further separated by the physical movement of the spindle poles themselves. Premature dissociation of the chromatids can lead to chromosome missegregation and aneuploidy in the daughter cells. Thus, the job of the metaphase checkpoint is to prevent this transition into anaphase until the chromosomes are properly attached, before the sister chromatids separate.
Spindle assembly checkpoint overview
The spindle assembly checkpoint (SAC) is an active signal produced by improperly attached kinetochores, which is conserved in all eukaryotes. The SAC stops the cell cycle by negatively regulating CDC20, thereby preventing the activation of the polyubiquitylation activities of anaphase promoting complex (APC). The proteins responsible for the SAC signal compose the mitotic checkpoint complex (MCC), which includes SAC proteins, MAD2/MAD3 (mitotic arrest deficient), BUB3 (budding uninhibited by benzimidazole), and CDC20.[52] Other proteins involved in the SAC include MAD1, BUB1, MPS1, and Aurora B. For higher eukaryotes, additional regulators of the SAC include constituents of the ROD-ZW10 complex, p31comet, MAPK, CDK1-cyclin-B, NEK2, and PLK1.[53]

Checkpoint activation
The SAC monitors the interaction between improperly connected kinetochores and spindle microtubules, and is maintained until kinetochores are properly attached to the spindle. During prometaphase, CDC20 and the SAC proteins concentrate at the kinetochores before attachment to the spindle assembly. These proteins keep the SAC activated until they are removed and the correct kinetochore-microtubule attachment is made. Even a single unattached kinetochore can maintain the spindle checkpoint.[52] After attachment of microtubule plus-ends and formation of kinetochore microtubules, MAD1 and MAD2 are depleted from the kinetochore assembly. Another regulator of checkpoint activation is kinetochore tension. When sister kinetochores are properly attached to opposite spindle poles, forces in the mitotic spindle generate tension at the kinetochores. Bi-oriented sister kinetochores stabilize the kinetochore-microtubule assembly whereas weak tension has a destabilizing effect. In response to incorrect kinetochore attachments such as syntelic attachment, where both kinetochores becomes attached to one spindle pole, the weak tension generated destabilizes the incorrect attachment and allows the kinetochore to reattach correctly to the spindle body. During this process, kinetochores that are attached to the mitotic spindle but that are not under tension trigger the spindle checkpoint. Aurora-B/Ipl1 kinase of the chromosomal passenger complex functions as the tensions sensor in improper kinetochore attachments. It detects and destabilizes incorrect attachments through control of the microtubule-severing KINI kinesin MCAK, the DASH complex, and the Ndc80/Hec1 complex[54] at the microtubule-kinetochore interface.[53] The Aurora-B/Ipl1 kinase is also critical in correcting merotelic attachments, where one kinetochore is simultaneously attached to both spindle poles. Merotelic attachments generate sufficient tension and are not detected by the SAC, and without correction, may result in chromosome mis-segregation due to slow chromatid migration speed. While microtubule attachment is independently required for SAC activation, it is unclear whether tension is an independent regulator of SAC, although it is clear that differing regulatory behaviors arise with tension.
Once activated, the spindle checkpoint blocks anaphase entry by inhibiting the anaphase-promoting complex via regulation of the activity of mitotic checkpoint complex. The mechanism of inhibition of APC by the mitotic checkpoint complex is poorly understood, although it is hypothesized that the MCC binds to APC as a pseudosubstrate using the KEN-box motif in BUBR1. At the same time that mitotic checkpoint complex is being activated, the centromere protein CENP-E activates BUBR1, which also blocks anaphase.[53]
Mitotic Checkpoint Complex formation
The mitotic checkpoint complex is composed of BUB3 together with MAD2 and MAD3 bound to Cdc20. MAD2 and MAD3 have distinct binding sites on CDC20, and act synergistically to inhibit APC/C. The MAD3 complex is composed of BUB3, which binds to Mad3 and BUB1B through the short linear motif known as the GLEBS motif. The exact order of attachments which must take place in order to form the MCC remains unknown. It is possible that Mad2-Cdc20 form a complex at the same time as BUBR1-BUB3-Cdc20 form another complex, and these two subcomplexes are consequently combined to form the mitotic checkpoint complex.[52] In human cells, binding of BUBR1 to CDC20 requires prior binding of MAD2 to CDC20, so it is possible that the MAD2-CDC20 subcomplex acts as an initiator for MCC formation. BUBR1 depletion leads only to a mild reduction in Mad2-Cdc20 levels while Mad2 is required for the binding of BubR1-Bub3 to Cdc20. Nevertheless BUBR1 is still required for checkpoint activation.[53]
The mechanism of formation for the MCC is unclear and there are competing theories for both kinetochore-dependent and kinetochore-independent formation. In support of the kinetochore-independent theory, MCC is detectable in S. cerevisiae cells in which core kinetocore assembly proteins have been mutated and cells in which the SAC has been deactivated, which suggests that the MCC could be assembled during mitosis without kinetochore localization. In one model, unattached prometaphase kinetochores can 'sensitize' APC to inhibition of MCC by recruiting the APC to kinetochores via a functioning SAC. Furthermore, depletions of various SAC proteins have revealed that MAD2 and BUBR1 depletions affect the timing of mitosis independently of kinetochores, while depletions of other SAC proteins result in a dysfunctional SAC without altering the duration of mitosis. Thus it is possible that the SAC functions through a two-stage timer where MAD2 and BUBR1 control the duration of mitosis in the first stage, which may be extended in the second stage if there are unattached kinetochores as well as other SAC proteins.[53] However, there are lines of evidence which are in disfavor of the kinetochore-independent assembly. MCC has yet to be found during interphase, while MCC does not form from its constituents in X. laevis meiosis II extracts without the addition of sperm of nuclei and nocodazole to prevent spindle assembly.
The leading model of MCC formation is the "MAD2-template model", which depends on the kinetochore dynamics of MAD2 to create the MCC. MAD1 localizes to unattached kinetochores while binding strongly to MAD2. The localization of MAD2 and BubR1 to the kinetochore may also be dependent on the Aurora B kinase.[55] Cells lacking Aurora B fail to arrest in metaphase even when chromosomes lack microtubule attachment.[56] Unattached kinetochores first bind to a MAD1-C-MAD2-p31comet complex and releases the p31comet through unknown mechanisms. The resulting MAD-C-MAD2 complex recruits the open conformer of Mad2 (O-Mad2) to the kinetochores. This O-Mad2 changes its conformation to closed Mad2 (C-Mad2) and binds Mad1. This Mad1/C-Mad2 complex is responsible for the recruitment of more O-Mad2 to the kinetochores, which changes its conformation to C-Mad2 and binds Cdc20 in an auto-amplification reaction. Since MAD1 and CDC20 both contain a similar MAD2-binding motif, the empty O-MAD2 conformation changes to C-MAD2 while binding to CDC20. This positive feedback loop is negatively regulated by p31comet, which competitively binds to C-MAD2 bound to either MAD1 or CDC20 and reduces further O-MAD2 binding to C-MAD2. Further control mechanisms may also exist, considering that p31comet is not present in lower eukaryotes. The 'template model' nomenclature is thus derived from the process where MAD1-C-MAD2 acts as a template for the formation of C-MAD2-CDC20 copies. This sequestration of Cdc20 is essential for maintaining the spindle checkpoint.[52]
Checkpoint deactivation
Several mechanisms exist to deactivate the SAC after correct bi-orientation of sister chromatids. Upon microtubule-kinetochore attachment, a mechanism of stripping via a dynein-dynein motor complex transports spindle checkpoint proteins away from the kinetochores.[53] The stripped proteins, which include MAD1, MAD2, MPS1, and CENP-F, are then redistributed to the spindle poles. The stripping process is highly dependent on undamaged microtubule structure as well as dynein motility along microtubules. As well as functioning as a regulator of the C-MAD2 positive feedback loop, p31comet also may act as a deactivator of the SAC. Unattached kinetochores temporarily inactivate p31comet, but attachment reactivates the protein and inhibits MAD2 activation, possibly by inhibitory phosphorylation. Another possible mechanism of SAC inactivation results from energy-dependent dissociation of the MAD2-CDC20 complex through non-degradative ubiquitylation of CDC20. Conversely, the de-ubiquitylating enzyme protectin is required to maintain the SAC. Thus, unattached kinetochores maintain the checkpoint by continuously recreating the MAD2-CDC20 subcomplex from its components. The SAC may also be deactivated by APC activation induced proteolysis. Since the SAC is not reactivated by the loss of sister-chromatid cohesion during anaphase, the proteolysis of cyclin B and inactivation of the CDK1-cyclin-B kinase also inhibits SAC activity. Degradation of MPS1 during anaphase prevents the reactivation of SAC after removal of sister-chromatid cohesion. After checkpoint deactivation and during the normal anaphase of the cell cycle, the anaphase promoting complex is activated through decreasing MCC activity. When this happens the enzyme complex polyubiquitinates the anaphase inhibitor securin. The ubiquitination and destruction of securin at the end of metaphase releases the active protease called separase. Separase cleaves the cohesion molecules that hold the sister chromatids together to activate anaphase.[57]
Spindle checkpoint defects and cancer
When the spindle checkpoint misfunctions, this can lead to chromosome missegregation, aneuploidy and even tumorigenesis.[53] Transformation occurs and is accelerated when maintenance of genomic integrity breaks down especially at the gross level of whole chromosomes or large portions of them. In fact, aneuploidy is the most common characteristic of human solid tumors and thus the spindle assembly checkpoint might be regarded as a possible target for anti-tumour therapy.[58] This is a much underappreciated fact since mutations in specific genes known as oncogenes or tumor suppressor are primarily thought to be behind genetic instability and tumorigenesis. Usually the various checkpoints in the cell cycle take care of genomic integrity via highly conserved redundant mechanisms that are important for maintaining cellular homeostasis and preventing tumorigenesis. Several spindle assembly checkpoint proteins act both as positive and negative regulators to ensure the proper chromosome segregation in each cell cycle preventing chromosome instability (CIN) also known as genome instability.
Genomic integrity is now appreciated at several levels where some tumors display instability manifested as base substitutions, insertions, and deletions, while the majority displays gains or losses of whole chromosomes.[59]
Due to the fact that alterations in mitotic regulatory proteins can lead to aneuploidy and this is a frequent event in cancer,[60] it was initially thought that these genes could be mutated in cancerous tissues.[61]
Mutated genes in cancers
In some cancers the genes that underlie the defects resulting in transformation are well characterized. In the hematological cancers such as multiple myeloma cytogenetic abnormalities are very common due to the inherent nature of DNA breaks needed for immunoglobulin gene rearrangement. However, defects in proteins such as MAD2 that function predominantly at the SAC also are characterized in multiple myeloma.[62] Most solid tumors are also predominantly aneuploid. For colorectal cancer, BUB1 and BUBR1 and amplification of STK15 are key regulators that have been implicated in the genomic instability resulting in cancer.[63] In breast cancer, the genetic form characterized by the BRCA-1 gene exhibits greater levels of genomic instability than sporadic forms. Experiments showed that BRCA-1 null mice have decreased expression of the key spindle checkpoint protein MAD2 .[64] For other cancers, more work is warranted to identify the causes of aneuploidy.
Other genes not traditionally associated with the SAC in cancer
Clearly variations in the physiological levels of these proteins (such as Mad2 or BubR1) are associated with aneuploidy and tumorigenesis, and this has been demonstrated using animal models.[65][66] However, recent studies indicate that what seems to happen is a more complicated scenario: aneuploidy would drive a high incidence of tumorigenesis only when alterations in the levels of specific mitotic checkpoint components (either reduction or overexpression) in tissues is also inducing other defects able to predispose them to tumors.[67] That is, defects such as an increase in DNA damage, chromosomal rearrangements, and/or a decreased incidence of cell death. For some mitotic checkpoint components, it is known that they are implicated in functions outside mitosis: nuclear import (Mad1), transcriptional repression (Bub3), and cell death, DNA damage response, aging, and megakaryopoiesis for BubR1. All this supports the conclusion that increase in tumorigenesis is associated with defects other than aneuploidy alone.[67]
Cancer-associated mutations affecting known checkpoint genes like BUB1 or BUBR1 are actually rare. However, several proteins implicated in cancer have intersections to spindle assembly networks. Key tumor suppressors such as p53 also play a role in the spindle checkpoint. Absence of p53, the most commonly mutated gene in human cancer, has a major effect on cell cycle checkpoint regulators and has been shown to act at the G1 checkpoint in the past, but now appears to be important in regulating the spindle checkpoint as well.[68] Another key aspect of cancer is inhibition of cell death or apoptosis. Survivin, a member of the inhibitor of apoptosis (IAP) family, is localized in pools at microtubules of the mitotic spindle near the centrosomes and at the kinetochores of metaphase chromosomes. Not only does survivin inhibit apoptosis to promote tumorigenesis, but it has been implicated (through experimental knockout mice) as an important regulator of chromosome segregation, and late stage mitosis similar to its role in more primitive organisms.[69]
Other aspects of the spindle assembly checkpoint such as kinetochore attachment, microtubule function, and sister chromatid cohesion are likely to be defective as well to cause aneuploidy. Cancer cells have been observed to divide in multiple directions by evading the spindle assembly checkpoint resulting in multipolar mitoses.[70] The multipolar metaphase-anaphase transition occurs through an incomplete separase cycle that results in frequent nondisjunction events which amplify aneuploidy in cancer cells.
SAC cancer therapies

Advances in this field have led to the introduction of development of some therapies targeted at spindle assembly defects. Older treatments such as vinca alkaloids and taxanes target microtubules that accompany mitotic spindle formation via disruption of microtubule dynamics which engage the SAC arresting the cell and eventually leading to its death.[71] taxol and Docetaxel both are still used in the treatment of breast cancer, ovarian cancer and other types of epithelial cancer. However, these treatments are often characterized by high rates of side effects and drug resistance.
Other targets within the network of regulators that influence the SAC are also being pursued; strong interest has shifted towards the aurora kinase proteins.[72] The kinase gene Aurora A when amplified acts as an oncogene overriding the SAC leading to abnormal initiation of anaphase and subsequent aneuploidy and also resistance to TAXOL .[73] Excitingly, a small molecule inhibitor of Aurora A has shown antitumor effects in an in vivo model suggesting that this might be a good target for further clinical development.[74] Aurora B inhibitors, which are also in clinical development lead to abnormal kinetochore to microtubule attachment and abrogate the mitotic checkpoint as well.[72] Survivin is also an attractive molecular target for clinical therapeutic development as it acts as a major node in a multitude of pathways, one of which is spindle formation and checkpoint control.[75] Even further approaches have included a look at inhibition of mitotic motor proteins like KSP. These inhibitors, which have recently entered clinical trials, cause mitotic arrest and by engaging the spindle assembly checkpoint and induce apoptosis.[76]
References
- ↑ Conlon, Ian; Raff, Martin (1999). "Size Control in Animal Development". Cell. 96 (2): 235–44. doi:10.1016/S0092-8674(00)80563-2. PMID 9988218.
- ↑ Nigg, Erich A.; Meraldi, Patrick; Lukas, Jiri; Fry, Andrew M.; Bartek, Jiri (1999). "Centrosome duplication in mammalian somatic cells requires E2F and Cdk2-cyclin A". Nature Cell Biology. 1 (2): 88–93. doi:10.1038/10054. PMID 10559879.
- ↑ Mayor, T; Meraldi, P; Stierhof, YD; Nigg, EA; Fry, AM (1999). "Protein kinases in control of the centrosome cycle". FEBS Letters. 452 (1–2): 92–5. doi:10.1016/S0014-5793(99)00534-7. PMID 10376685.
- ↑ Nicklas, R. B. (1997). "How Cells Get the Right Chromosomes". Science. 275 (5300): 632–7. doi:10.1126/science.275.5300.632. PMID 9005842.
- ↑ Lončarek, Jadranka; Kisurina-Evgenieva, Olga; Vinogradova, Tatiana; Hergert, Polla; La Terra, Sabrina; Kapoor, Tarun M.; Khodjakov, Alexey (2007). "The centromere geometry essential for keeping mitosis error free is controlled by spindle forces". Nature. 450 (7170): 745–9. doi:10.1038/nature06344. PMC 2586812
 . PMID 18046416.
. PMID 18046416. - ↑ Dewar, Hilary; Tanaka, Kozo; Nasmyth, Kim; Tanaka, Tomoyuki U. (2004). "Tension between two kinetochores suffices for their bi-orientation on the mitotic spindle". Nature. 428 (6978): 93–7. doi:10.1038/nature02328. PMID 14961024.
- ↑ Cimini, Daniela; Wan, Xiaohu; Hirel, Christophe B.; Salmon, E.D. (2006). "Aurora Kinase Promotes Turnover of Kinetochore Microtubules to Reduce Chromosome Segregation Errors". Current Biology. 16 (17): 1711–8. doi:10.1016/j.cub.2006.07.022. PMID 16950108.
- ↑ Gautschi, O.; Heighway, J.; Mack, P. C.; Purnell, P. R.; Lara, P. N.; Gandara, D. R. (2008). "Aurora Kinases as Anticancer Drug Targets". Clinical Cancer Research. 14 (6): 1639–48. doi:10.1158/1078-0432.CCR-07-2179. PMID 18347165.
- 1 2 Zirkle, R. E. (1970). "Ultraviolet-Microbeam Irradiation of Newt-Cell Cytoplasm: Spindle Destruction, False Anaphase, and Delay of True Anaphase". Radiation Research. 41 (3): 516–37. doi:10.2307/3572841. JSTOR 3572841. PMID 5438206.
- ↑ Rieder, CL; Palazzo, RE (1992). "Colcemid and the mitotic cycle". Journal of Cell Science. 102 (3): 387–92. PMID 1506421.
- ↑ Burke, Daniel J.; Stukenberg, P. Todd (2008). "Linking Kinetochore-Microtubule Binding to the Spindle Checkpoint". Developmental Cell. 14 (4): 474–9. doi:10.1016/j.devcel.2008.03.015. PMC 2696048
 . PMID 18410725.
. PMID 18410725. - 1 2 Li, Rong; Murray, Andrew W. (1991). "Feedback control of mitosis in budding yeast". Cell. 66 (3): 519–31. doi:10.1016/0092-8674(81)90015-5. PMID 1651172.
- ↑ Hoyt, M.Andrew; Totis, Laura; Roberts, B.Tibor (1991). "S. cerevisiae genes required for cell cycle arrest in response to loss of microtubule function". Cell. 66 (3): 507–17. doi:10.1016/0092-8674(81)90014-3. PMID 1651171.
- ↑ Neff, MW; Burke, DJ (1992). "A delay in the Saccharomyces cerevisiae cell cycle that is induced by a dicentric chromosome and dependent upon mitotic checkpoints". Molecular and Cellular Biology. 12 (9): 3857–64. PMC 360258
 . PMID 1324407.
. PMID 1324407. - ↑ Wells, W. A.; Murray, AW (1996). "Aberrantly segregating centromeres activate the spindle assembly checkpoint in budding yeast". The Journal of Cell Biology. 133 (1): 75–84. doi:10.1083/jcb.133.1.75. PMC 2120768
 . PMID 8601615.
. PMID 8601615. - ↑ Hardwick, K. G.; Weiss, E.; Luca, F. C.; Winey, M.; Murray, A. W. (1996). "Activation of the Budding Yeast Spindle Assembly Checkpoint Without Mitotic Spindle Disruption". Science. 273 (5277): 953–6. doi:10.1126/science.273.5277.953. PMID 8688079.
- ↑ Wang, Y; Burke, DJ (1995). "Checkpoint genes required to delay cell division in response to nocodazole respond to impaired kinetochore function in the yeast Saccharomyces cerevisiae". Molecular and Cellular Biology. 15 (12): 6838–44. PMC 230938
 . PMID 8524250.
. PMID 8524250. - ↑ Spencer, F.; Hieter, P. (1992). "Centromere DNA Mutations Induce a Mitotic Delay in Saccharomyces cerevisiae". Proceedings of the National Academy of Sciences of the United States of America. 89 (19): 8908–8912. doi:10.1073/pnas.89.19.8908. JSTOR 2360300. PMC 50033
 . PMID 1409584.
. PMID 1409584. - ↑ Hardwick, KG; Li, R; Mistrot, C; Chen, RH; Dann, P; Rudner, A; Murray, AW (1999). "Lesions in many different spindle components activate the spindle checkpoint in the budding yeast Saccharomyces cerevisiae". Genetics. 152 (2): 509–18. PMC 1460633
 . PMID 10353895.
. PMID 10353895. - ↑ McIntosh, JR (1991). "Structural and mechanical control of mitotic progression". Cold Spring Harbor symposia on quantitative biology. 56: 613–9. doi:10.1101/sqb.1991.056.01.070. PMID 1819511.
- 1 2 Rieder, C. L.; Cole, RW; Khodjakov, A; Sluder, G (1995). "The checkpoint delaying anaphase in response to chromosome monoorientation is mediated by an inhibitory signal produced by unattached kinetochores". The Journal of Cell Biology. 130 (4): 941–8. doi:10.1083/jcb.130.4.941. PMC 2199954
 . PMID 7642709.
. PMID 7642709. - ↑ Li, X; Nicklas, RB (1997). "Tension-sensitive kinetochore phosphorylation and the chromosome distribution checkpoint in praying mantid spermatocytes". Journal of Cell Science. 110 (5): 537–45. PMID 9092936.
- ↑ Rieder, C. L.; Khodjakov, A.; Paliulis, L. V.; Fortier, T. M.; Cole, R. W.; Sluder, G. (1997). "Mitosis in vertebrate somatic cells with two spindles: Implications for the metaphase/anaphase transition checkpoint and cleavage". Proceedings of the National Academy of Sciences. 94 (10): 5107–12. doi:10.1073/pnas.94.10.5107.
- 1 2 Hirano, Tatsuya (2000). "CHROMOSOMECOHESION, CONDENSATION, ANDSEPARATION". Annual Review of Biochemistry. 69: 115–44. doi:10.1146/annurev.biochem.69.1.115. PMID 10966455.
- ↑ Tanaka, K.; Hao, Z; Kai, M; Okayama, H (2001). "Establishment and maintenance of sister chromatid cohesion in fission yeast by a unique mechanism". The EMBO Journal. 20 (20): 5779–90. doi:10.1093/emboj/20.20.5779. PMC 125673
 . PMID 11598020.
. PMID 11598020. - ↑ Gonzalez, Cayetano; Jimenez, Jose Casal; Ripoll, Pedro; Sunkel, Claudio E. (1991). "The spindle is required for the process of sister chromatid separation in Drosophila neuroblasts". Experimental Cell Research. 192 (1): 10–5. doi:10.1016/0014-4827(91)90150-S. PMID 1898588.
- ↑ Losada, Ana; Hirano, Tatsuya (2001). "Shaping the metaphase chromosome: coordination of cohesion and condensation". BioEssays. 23 (10): 924–35. doi:10.1002/bies.1133. PMID 11598959.
- ↑ Bernard, P. (2001). "Requirement of Heterochromatin for Cohesion at Centromeres". Science. 294 (5551): 2539–542. doi:10.1126/science.1064027.
- ↑ Nonaka, Nobuhiro; Kitajima, Tomoya; Yokobayashi, Shihori; Xiao, Guoping; Yamamoto, Masayuki; Grewal, Shiv I. S.; Watanabe, Yoshinori (2001). "Recruitment of cohesin to heterochromatic regions by Swi6/HP1 in fission yeast". Nature Cell Biology. 4 (1): 89–93. doi:10.1038/ncb739. PMID 11780129.
- ↑ Hall, I. M.; Noma, K.-i.; Grewal, S. I. S. (2002). "RNA interference machinery regulates chromosome dynamics during mitosis and meiosis in fission yeast". Proceedings of the National Academy of Sciences. 100: 193–8. doi:10.1073/pnas.232688099.
- ↑ Fukagawa, Tatsuo; Nogami, Masahiro; Yoshikawa, Mitsuko; Ikeno, Masashi; Okazaki, Tuneko; Takami, Yasunari; Nakayama, Tatsuo; Oshimura, Mitsuo (2004). "Dicer is essential for formation of the heterochromatin structure in vertebrate cells". Nature Cell Biology. 6 (8): 784–91. doi:10.1038/ncb1155. PMID 15247924.
- ↑ Weber, Stewart A.; Gerton, Jennifer L.; Polancic, Joan E.; Derisi, Joseph L.; Koshland, Douglas; Megee, Paul C. (2004). "The Kinetochore Is an Enhancer of Pericentric Cohesin Binding". PLoS Biology. 2 (9): e260. doi:10.1371/journal.pbio.0020260. PMC 490027
 . PMID 15309047.
. PMID 15309047. 
- ↑ Prasanth, Supriya G; Prasanth, Kannanganattu V; Siddiqui, Khalid; Spector, David L; Stillman, Bruce (2004). "Human Orc2 localizes to centrosomes, centromeres and heterochromatin during chromosome inheritance". The EMBO Journal. 23 (13): 2651–63. doi:10.1038/sj.emboj.7600255. PMC 449767
 . PMID 15215892.
. PMID 15215892. - ↑ Shimada, Kenji; Gasser, Susan M. (2007). "The Origin Recognition Complex Functions in Sister-Chromatid Cohesion in Saccharomyces cerevisiae". Cell. 128 (1): 85–99. doi:10.1016/j.cell.2006.11.045. PMID 17218257.
- ↑ Kato, H; Matsunaga, F; Miyazaki, S; Yin, L; D'urso, G; Tanaka, K; Murakami, Y (2008). "Schizosaccharomyces pombe Orc5 plays multiple roles in the maintenance of genome stability throughout the cell cycle". Cell cycle. 7 (8): 1085–96. doi:10.4161/cc.7.8.5710. PMID 18414064.
- ↑ Hauf, Silke; Watanabe, Yoshinori (2004). "Kinetochore Orientation in Mitosis and Meiosis". Cell. 119 (3): 317–27. doi:10.1016/j.cell.2004.10.014. PMID 15507205.
- ↑ Sonoda, Eiichiro; Matsusaka, Takahiro; Morrison, Ciaran; Vagnarelli, Paola; Hoshi, Osamu; Ushiki, Tatsuo; Nojima, Kuniharu; Fukagawa, Tatsuo; et al. (2001). "Scc1/Rad21/Mcd1 Is Required for Sister Chromatid Cohesion and Kinetochore Function in Vertebrate Cells". Developmental Cell. 1 (6): 759–70. doi:10.1016/S1534-5807(01)00088-0. PMID 11740938.
- ↑ Vass, Sharron; Cotterill, Sue; Valdeolmillos, Ana M.; Barbero, José L.; Lin, Enmoore; Warren, William D.; Heck, Margarete M.S. (2003). "Depletion of Drad21/Scc1 in Drosophila Cells Leads to Instability of the Cohesin Complex and Disruption of Mitotic Progression". Current Biology. 13 (3): 208–18. doi:10.1016/S0960-9822(03)00047-2. PMID 12573216.
- ↑ Haering, Christian H.; Löwe, Jan; Hochwagen, Andreas; Nasmyth, Kim (2002). "Molecular Architecture of SMC Proteins and the Yeast Cohesin Complex". Molecular Cell. 9 (4): 773–88. doi:10.1016/S1097-2765(02)00515-4. PMID 11983169.
- ↑ Hirano, T. (1999). "SMC-mediated chromosome mechanics: a conserved scheme from bacteria to vertebrates?". Genes & Development. 13: 11–9. doi:10.1101/gad.13.1.11.
- ↑ Ciosk, Rafal; Zachariae, Wolfgang; Michaelis, Christine; Shevchenko, Andrej; Mann, Matthias; Nasmyth, Kim (1998). "An ESP1/PDS1 Complex Regulates Loss of Sister Chromatid Cohesion at the Metaphase to Anaphase Transition in Yeast". Cell. 93 (6): 1067–76. doi:10.1016/S0092-8674(00)81211-8. PMID 9635435.
- ↑ Alexandru, Gabriela; Uhlmann, Frank; Mechtler, Karl; Poupart, Marc-André; Nasmyth, Kim (2001). "Phosphorylation of the Cohesin Subunit Scc1 by Polo/Cdc5 Kinase Regulates Sister Chromatid Separation in Yeast". Cell. 105 (4): 459–72. doi:10.1016/S0092-8674(01)00362-2. PMID 11371343.
- ↑ Leismann, O. (2000). "Degradation of Drosophila PIM regulates sister chromatid separation during mitosis". Genes & Development. 14 (17): 2192–205. doi:10.1101/gad.176700.
- ↑ Zur, A.; Brandeis, M (2001). "Securin degradation is mediated by fzy and fzr, and is required for complete chromatid separation but not for cytokinesis". The EMBO Journal. 20 (4): 792–801. doi:10.1093/emboj/20.4.792. PMC 145417
 . PMID 11179223.
. PMID 11179223. - ↑ Sumara, I.; Vorlaufer, E.; Gieffers, C.; Peters, B. H.; Peters, J.-M. (2000). "Characterization of Vertebrate Cohesin Complexes and Their Regulation in Prophase". The Journal of Cell Biology. 151 (4): 749–62. doi:10.1083/jcb.151.4.749. PMC 2169443
 . PMID 11076961.
. PMID 11076961. - ↑ Losada, A.; Yokochi, T.; Kobayashi, R.; Hirano, T. (2000). "Identification and Characterization of Sa/Scc3p Subunits in the Xenopus and Human Cohesin Complexes". The Journal of Cell Biology. 150 (3): 405–16. doi:10.1083/jcb.150.3.405. PMC 2175199
 . PMID 10931856.
. PMID 10931856. - ↑ Giménez-Abián, Juan F; Sumara, Izabela; Hirota, Toru; Hauf, Silke; Gerlich, Daniel; De La Torre, Consuelo; Ellenberg, Jan; Peters, Jan-Michael (2004). "Regulation of Sister Chromatid Cohesion between Chromosome Arms". Current Biology. 14 (13): 1187–93. doi:10.1016/j.cub.2004.06.052. PMID 15242616.
- ↑ Paliulis, Leocadia V.; Nicklas, R.Bruce (2004). "Micromanipulation of Chromosomes Reveals that Cohesion Release during Cell Division Is Gradual and Does Not Require Tension". Current Biology. 14 (23): 2124–9. doi:10.1016/j.cub.2004.11.052. PMID 15589155.
- ↑ Nakajima, M.; Kumada, K.; Hatakeyama, K.; Noda, T.; Peters, J.-M.; Hirota, T. (2007). "The complete removal of cohesin from chromosome arms depends on separase". Journal of Cell Science. 120 (23): 4188. doi:10.1242/jcs.011528.
- ↑ McGuinness, Barry E.; Hirota, Toru; Kudo, Nobuaki R.; Peters, Jan-Michael; Nasmyth, Kim (2005). "Shugoshin Prevents Dissociation of Cohesin from Centromeres During Mitosis in Vertebrate Cells". PLoS Biology. 3 (3): e86. doi:10.1371/journal.pbio.0030086. PMC 1054882
 . PMID 15737064.
. PMID 15737064. 
- ↑ Salic, Adrian; Waters, Jennifer C.; Mitchison, Timothy J. (2004). "Vertebrate Shugoshin Links Sister Centromere Cohesion and Kinetochore Microtubule Stability in Mitosis". Cell. 118 (5): 567–78. doi:10.1016/j.cell.2004.08.016. PMID 15339662.
- 1 2 3 4 De Antoni, Anna; Pearson, Chad G.; Cimini, Daniela; Canman, Julie C.; Sala, Valeria; Nezi, Luigi; Mapelli, Marina; Sironi, Lucia; et al. (2005). "The Mad1/Mad2 Complex as a Template for Mad2 Activation in the Spindle Assembly Checkpoint". Current Biology. 15 (3): 214–25. doi:10.1016/j.cub.2005.01.038. PMID 15694304.
- 1 2 3 4 5 6 7 Musacchio, Andrea; Edward D. Salmon (May 2007). "The spindle-assembly checkpoint in space and time". Nat Rev Mol Cell Biol. 8 (5): 379–393. doi:10.1038/nrm2163. ISSN 1471-0072. PMID 17426725.
- ↑ Martin-Lluesma, S.; Stucke, VM; Nigg, EA (2002). "Role of Hec1 in Spindle Checkpoint Signaling and Kinetochore Recruitment of Mad1/Mad2". Science. 297 (5590): 2267–70. doi:10.1126/science.1075596. PMID 12351790.
- ↑ Lens, Susanne M.A.; Rob M.F. Wolthuis; Rob Klompmaker; Jos Kauw; Reuven Agami; Thijn Brummelkamp; Geert Kops; René H. Medema (2003-06-16). "Survivin is required for a sustained spindle checkpoint arrest in response to lack of tension". The EMBO Journal. 22 (12): 2934–2947. doi:10.1093/emboj/cdg307. ISSN 0261-4189. PMC 162159
 . PMID 12805209.
. PMID 12805209. - ↑ Hauf, S; Cole, R. W.; Laterra, S; Zimmer, C; Schnapp, G; Walter, R; Heckel, A; Van Meel, J; Rieder, C. L.; Peters, J. M. (2003). "The small molecule Hesperadin reveals a role for Aurora B in correcting kinetochore–microtubule attachment and in maintaining the spindle assembly checkpoint". The Journal of Cell Biology. 161 (2): 281–294. doi:10.1083/jcb.200208092. PMC 2172906
 . PMID 12707311.
. PMID 12707311. - ↑ Morgan, David O. (2006-09-06). The Cell Cycle: Principles of Control (Primers in Biology) (1 ed.). New Science Press, Ltd. ISBN 0-87893-508-8.
- ↑ Kops, Geert J.P.L.; Weaver, Beth A. A.; Cleveland, Don W. (2005). "On the road to cancer: aneuploidy and the mitotic checkpoint". Nature Reviews Cancer. 5 (10): 773–785. doi:10.1038/nrc1714. PMID 16195750.
- ↑ Lengauer, Christoph; Kinzler, Kenneth W.; Vogelstein, Bert (1998). "Genetic instabilities in human cancers". Nature. 396 (6712): 643–649. doi:10.1038/25292. PMID 9872311.
- ↑ Weaver, Beth AA; Cleveland, Don W (2006). "Does aneuploidy cause cancer?". Current Opinion in Cell Biology. 18 (6): 658–67. doi:10.1016/j.ceb.2006.10.002. PMID 17046232.
- ↑ Lengauer, Christoph; Cahill, Daniel P.; Yu, Jian; Riggins, Gregory J.; Willson, James K. V.; Markowitz, Sanford D.; Kinzler, Kenneth W.; Vogelstein, Bert (1998). "Mutations of mitotic checkpoint genes in human cancers". Nature. 392 (6673): 300–3. doi:10.1038/32688. PMID 9521327.
- ↑ Diaz-Rodriguez, Elena; Álvarez-Fernández, Stela; Chen, Xi; Paiva, Bruno; López-Pérez, Ricardo; García-Hernández, Juan Luis; San Miguel, Jesús F.; Pandiella, Atanasio (2011). "Deficient Spindle Assembly Checkpoint in Multiple Myeloma". PLoS ONE. 6 (11): e27583. doi:10.1371/journal.pone.0027583. PMC 3223182
 . PMID 22132115.
. PMID 22132115. 
- ↑ Grady, William M. (2004). "Genomic instability and colon cancer". Cancer and Metastasis Review. 23 (1–2): 11–27. doi:10.1023/A:1025861527711.
- ↑ Wang, Rui-Hong; Hongtao, Yu; Deng, Chu-Xia (2004). "A requirement for breast-cancer associated gene (BRCA1) in the spindle checkpoint". PNAS. 101 (49): 17108–17113. doi:10.1073/pnas.0407585101. PMC 535394
 . PMID 15563594.
. PMID 15563594. - ↑ Sotillo, Rocío; Hernando, Eva; Díaz-Rodríguez, Elena; Teruya-Feldstein, Julie; Cordón-Cardo, Carlos; Lowe, Scott W.; Benezra, Robert (2007). "Mad2 Overexpression Promotes Aneuploidy and Tumorigenesis in Mice". Cancer Cell. 11 (1): 9–23. doi:10.1016/j.ccr.2006.10.019. PMC 1850996
 . PMID 17189715.
. PMID 17189715. - ↑ Yamamoto, Y; Matsuyama, H; Chochi, Y; Okuda, M; Kawauchi, S; Inoue, R; Furuya, T; Oga, A; et al. (2007). "Overexpression of BUBR1 is associated with chromosomal instability in bladder cancer". Cancer Genetics and Cytogenetics. 174 (1): 42–7. doi:10.1016/j.cancergencyto.2006.11.012. PMID 17350465.
- 1 2 Weaver, B. A.A.; Cleveland, D. W. (2009). "The role of aneuploidy in promoting and suppressing tumors". The Journal of Cell Biology. 185 (6): 935–7. doi:10.1083/jcb.200905098. PMC 2711620
 . PMID 19528293.
. PMID 19528293. - ↑ Cross, Shawn M.; Sanchez, Carissa A; Morgan, Catherine A.; Schimke, Melana K.; Reid, Brian J. (1995). "A p53-dependant mouse spindle checkpoint". Science. 3 (5202): 1353–1356. doi:10.1126/science.7871434.
- ↑ Altieri, Dario C. (2001). "The molecular basis and potential role of survivin in cancer diagnosis and therapy". Trends in Molecular Medicine. 7 (12): 542–547. doi:10.1016/S1471-4914(01)02243-2. PMID 11733216.
- ↑ Gisselsson, David; Hakanson, Ulf; Stoller, Patrick; Marti, Dominik; Jin, Yuesheng; Rosengren, Anders H.; Stewénius, Ylva; Kahl, Fredrik; Panagopoulos, Ioannis (2008). "When the Genome Plays Dice: Circumvention of the Spindle Assembly Checkpoint and Near-Random Chromosome Segregation in Multipolar Cancer Cell Mitoses". PLoS ONE. 3 (4): e1871. doi:10.1371/journal.pone.0001871. PMC 2289843
 . PMID 18392149.
. PMID 18392149. 
- ↑ Zhou, Jun; Giannakakou, Paraskevi (2005). "Targeting Microtubules for Cancer Chemotherapy". Current Medicinal Chemistry - Anti-Cancer Agents. 5 (1): 65–71. doi:10.2174/1568011053352569. PMID 15720262.
- 1 2 Carvajal, Richard D.; Tse, Archie; Schwartz, Gary K. (2006). "Aurora Kinases: New Targets for Cancer Therapy". Clinical Cancer Research. 12 (23): 6869–75. doi:10.1158/1078-0432.CCR-06-1405. PMID 17145803.
- ↑ Anand, Shubha; Penrhyn-Lowe, Sue; Venkitaraman, Ashok R. (2003). "Aurora-A amplification overrides the mitotic spindle assembly checkpoint, inducing resistance to Taxol". Cancer Cell. 3 (1): 51–62. doi:10.1016/S1535-6108(02)00235-0. PMID 12559175.
- ↑ Harrington, E. A.; Bebbington, D; Moore, J; Rasmussen, R. K.; Ajose-Adeogun, A. O.; Nakayama, T; Graham, J. A.; Demur, C; Hercend, T; Diu-Hercend, A; Su, M; Golec, J. M.; Miller, K. M.; et al. (2004). "VX-680, a potent and selective small-molecule inhibitor of the Aurora kinases, suppresses tumor growth in vivo". Nature Medicine. 10 (3): 262–267. doi:10.1038/nm1003. PMID 14981513.
- ↑ Altieri, Dario C. (2008). "Survivin, cancer networks and pathway-directed drug discovery". Nature Reviews Cancer. 8 (1): 61–70. doi:10.1038/nrc2293. PMID 18075512.
- ↑ Tao, Weikang; et al. (2005). "Induction of apoptosis by an inhibitor of the mitotic kinesin KSP requires both activation of the spindle assembly checkpoint and mitotic slippage". Cancer Cell. 8 (1): 49–59. doi:10.1016/j.ccr.2005.06.003. PMID 16023598.
Further reading
- Larsen, N. A.; Al-Bassam, J.; Wei, R. R.; Harrison, S. C. (2007). "Structural analysis of Bub3 interactions in the mitotic spindle checkpoint". Proceedings of the National Academy of Sciences. 104 (4): 1201–6. doi:10.1073/pnas.0610358104. PMC 1770893
 . PMID 17227844.
. PMID 17227844. - Wang, X.; Babu, JR; Harden, JM; Jablonski, SA; Gazi, MH; Lingle, WL; De Groen, PC; Yen, TJ; Van Deursen, JM (2001). "The Mitotic Checkpoint Protein hBUB3 and the mRNA Export Factor hRAE1 Interact with GLE2p-binding Sequence (GLEBS)-containing Proteins". Journal of Biological Chemistry. 276 (28): 26559–67. doi:10.1074/jbc.M101083200. PMID 11352911.
- Kitagawa, Risa; Rose, Ann M. (1999). "Components of the spindle-assembly checkpoint are essential in Caenorhabditis elegans". Nature Cell Biology. 1 (8): 514–21. doi:10.1038/70309. PMID 10587648.
External links
- Ted Salmon's lab: dividing cells movies.
- Andrea Musacchio's lab: spindle checkpoint schemes.
- http://www.uniprot.org/uniprot/O60566