Evolution of the wolf
_Wolf_Cranium.png)
The evolution of the wolf occurred over a geologic time scale of 800 thousand years, transforming the first Middle Pleistocene wolf specimen that is recognized as being morphologically similar to Canis lupus into today's dog, dingo and gray wolf. Ecological factors including habitat type, climate, prey specialization and predatory competition has greatly influenced the wolf's genetic population structure and cranio-dental plasticity. Wolves went through a population bottleneck 20,000 years before present (YBP), which indicates that many wolf populations had gone extinct at a time that coincided with the Last Glacial Maximum and the expansion of modern humans worldwide with their technology for capturing large game. The domestic dog is the most widely abundant large carnivore and a descendant from one of those now-extinct wolf populations. Today, the wolf is represented by the many extant subspecies of Canis lupus, which includes the dog and dingo.
Fossil record
- Further information: Evolution of Canis, and Canid relationships
The fossil record for ancient vertebrates is composed of rarely occurring fragments from which it is often impossible to obtain genetic material. Researchers are limited to morphologic analysis but it is difficult to estimate the intra-species and inter-species variations and relationships that existed between specimens across time and place. Some observations are debated by researchers who do not always agree, and hypotheses that are supported by some authors are challenged by others.[2]
There is general agreement on the most ancient record, which shows that Feliforms and Caniforms emerged within the super-family Carnivoramorpha 43 million years before present (YBP).[3] The caniforms included the fox-like Leptocyon genus whose various species existed from 34 million YBP before branching 11.9 million YBP into vulpes (foxes) and canini (canines). The jackal-sized Eucyon existed in North America from 10 million YBP and by the Early Pliocene about 6–5 million YBP the coyote-like Eucyon davisi[4] invaded Eurasia. In North America it gave rise to early Canis which first appeared in the Miocene (6 million YBP) in south-western US and Mexico. By 5 million YBP the larger Canis lepophagus appeared in the same region.[5]:p58
The canids that had immigranted from North America to Eurasia – Eucyon, Vulpes, and Nyctereutes – were small to medium-sized predators during the Late Miocene and Early Pliocene but they were not the top predators. The position of the canids would change with the arrival of Canis to become a dominant predator across the Holarctic. The wolf-sized C. chihilensis appeared in northern China in the Mid-Pliocene around 4–3 million YBP. This was followed by an explosion of Canis evolution across Eurasia in the Early Pleistocene around 1.8 million YBP in what is commonly referred to as the Wolf event. It is associated with the formation of the Mammoth steppe and continental glaciation. Canis spread to Europe in the forms of C. arnensis, C. eutruscus, and C. falconeri.[5]:p148
The fossil record is incomplete but it is likely that wolves arose from a population of small, early canids.[6]:p241 Morphological evidence[6]:p239[7] and genetic evidence[8] both suggest that wolves evolved during the Pliocene and Early Pleistocene eras from the same lineage that also led to the coyote,[6]:p239 with fossil specimens indicating that the coyote and the wolf diverged from a common ancestor 1.5 million years ago.[6]:p240[7] The ancestor of the jackal and the other extant members of the Canis genus had split from the lineage before this time.[6]:p240
After this separation from a common ancestor the species that were believed to be involved in the further evolution of the wolf and coyote - and the beliefs of some paleontologists - diverged.[6]:p240 A number of researchers believed that the lines of C. priscolatrans, C. etruscus, C. rufus and C. lupus were components involved in some way that lead to the modern wolf and coyote.[6]:p240[9][10][11][12][13][14]
| Wolf evolution | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Proposed evolution and branching of genus Canis towards the wolf.[6]:p240 |
Canis lepophagus
Canis lepophagus lived in the early Pliocene in North America.[5] Kurten proposed that the Blancan C. lepophagus[15] derived from smaller Miocene Canis species in North America. It then became widespread across Eurasia where it was either identical to, or closely related with, C. arnensis of Europe.[6]:p241[16][17]
Johnston describes C. lepophagus as having a more slender skull and skeleton than in the modern coyote.[18]:385 Nowak found that the early populations had small, delicate and narrowly proportioned skulls that resemble small coyotes and appear to be ancestral to C. latrans.[6]:p241 Johnson noted that some specimens found in Cita Canyon, Texas had larger, broader skulls,[18] and along with other fragments Nowak suggested that these were evolving into wolves.[6]:p241[7]
Tedford disagreed with previous authors and found that its cranio-dental morphology lacked some characteristics that are shared by C. lupus and C. latrans, and therefore there was not a close relationship but it did suggest C. lepophagus was the ancestor of both wolves and coyotes.[19]:p119
Canis priscolatrans
Canis priscolatrans lived in the late Pliocene-Early Pleistocene in North America.[7] The first definite wolf appeared in the Late Blancan/Early Irvingtonian,[6]:p240[7][20] and named C. priscolatrans that was either very close to[16][17] or a synonym for Canis edwardii.[6]:p241[7]:82[21][22] It resembled C. rufus in cranial size and proportions but with more complex dentition.[6]:p241 However, there are no fossils of C. rufus until the Late Rancholabrean.[6]:p242
Kurten was uncertain if C. priscolatrans derived from C. lepophagus and C. arnensis,[17] but believed that C. priscolatrans was a population of large coyotes that were ancestral to Rancholabrean and recent C. latrans. He noted that C. arnensis of Europe showed striking similarities to C. priscolatrans, and they could represent what once was a holarctic population of coyotes.[16]:p27 Nowak disagreed, and believed that C. priscolatrans was a counterpart to the European C. etruscus.[7] Kurten later proposed that both C. priscolatrans and C. etruscus were part of a group which led to C. lupus but was not sure if they evolved separately from C. lepophagus or a possible common ancestor that was derived from C. lepophagus.[17]
The remains of the larger coyote-like Canis edwardii have been found in the later Pliocene in the south-western USA along with C. lepophagus, which indicates a descent.[5]:p60 Tedford recognised C. edwardii[23] and found that the cranio-dental morphology of C. priscolatrans fell inside that of C. edwardii such that the species name C. priscolatrans was doubtful (nomen dubium).[19]:p131
Canis ambrusteri
The North American wolves became larger, with tooth specimens indicating that C. priscolatrans diverged into the large wolf C. ambrusteri.[6]:p242[24] during the Middle Pleistocene in North America.[7] Martin disagreed, and believed that C. ambrusteri[25] was C. lupus.[12] Nowak disagreed with Martin and proposed that C. ambrusteri was not related to C. lupus but C. priscolatrans, which then gave rise to C. dirus. Tedford proposed that the South American C. gezi and C. nehringi share dental and cranial similarities developed for hypercarnivory, suggesting C. ambrusteri was the common ancestor of C. gezi, C. nehringi and C. dirus.[19]:148
Canis dirus
Canis dirus lived in the late Pleistocene to early Holocene in North and South America.[26] C. dirus was the largest of all Canis species.[5]:52 Goulet disagreed with Nowak and agreed with Martin, and proposed that the dire wolf C. dirus[27] was a variant of C. lupus.[28] Nowak, Kurten and Berta disagreed with Goulet and proposed that C. dirus was not derived from C. lupus.[7][17][29] The three noted paleontologists Xiaoming Wang, R. H. Tedford and R. M. Nowak have all proposed that C. dirus had evolved from C. ambrusteri,[5]:p52[19]:181 with Nowak stating that there were specimens from Cumberland Cave, Maryland that indicated C. ambrusteri diverging into C. dirus.[6]:p243[30] The two taxa share a number of characteristics (synapomorphy), which suggests an origin of C. dirus in the late Irvingtonian in the open terrain in the midcontinent, and then later expanding eastward and displacing its ancestor C. ambrusteri.[19]:181
Canis mosbachensis
Canis mosbachensis lived in the middle to late Pleistocene in Eurasia.[31] The holotype of the Mosbach wolf C. mosbachensis (Soergel, 1925)[32] was found in Jockgrim, (Germany). In 2010, a study found that the diversity of the Canis group decreased by the end of the Early Pleistocene to Middle Pleistocene and was limited in Eurasia to the small wolves of the C. mosbachensis–C. variabilis group that were a comparable size to the extant Indian wolf (Canis lupus pallipes), and the large hypercarnivorous Canis (Xenocyon) lycaonoides that was comparable in size to extant northern gray wolves.[33] C. mosbachensis occurred between C. etruscus in the Early Pleistocene and the modern C. lupus.[6]:p242 C. mosbachensis was smaller than most North American wolf populations and smaller than C. rufus,[6]:p242[34] and has been described by Kurten as being similar in size to Canis papilles.[6]:p242[31] As wolves continue to evolve they become bigger. It was still living in Europe at a time coinciding with the North American Early Rancholabrean and had grown in size by the Late Rancholabrean. Nowak proposed that C. mosbachensis was the ancestor of Eurasian and North American wolves, and that one population of C. mosbachensis invaded North America where it became isolated by the later glaciation and there gave rise to C. rufus. Another population of C. mosbachensis remained in Eurasia and evolved into C. lupus, from where it invaded North America.[6]:p242
Tedford compared C. mosbachensis (which was once distributed from Western Europe to Khazakhstan) with C. variabilis (which was once distributed from Khazakhstan to China) as they both existed in the Middle Pleistocene across mid-latitude Eurasia. The only difference was that C. variabilis had "nasal bones that terminate at or anterior to the most posterior position of the frontal-maxillary suture", and therefore these two taxa could represent variation in the one geographically widespread mid-Pleistocene wolf.[19]:p181
The phylogenetic descent of the extant wolf C. lupus from C. etruscus through C. mosbachensis is widely accepted.[2][31][33][35][36][37][38][39][40][41][42][43][44] Lumley considered C. mosbachensis to be a subspecies of the gray wolf and proposed a name of C. lupus mosbachensis.[45] However, other researchers propose that they can see no clear anatomical relationship between C. mosbachensis and C. etruscus and that it is more similar to C. arnensis,[46][47][48] and that it exhibits a size and dentition more similar to an omnivorous jackal.[48]
Canis chihliensis
| Wolf evolution - alternate proposal | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Proposed evolution and branching from Eucyon towards the wolf.[5]:p148[19]:p181 |
Wang and Tedford proposed that the genus Canis was the descendant of the coyote-like Eucyon davisi, and its remains first appeared in the Miocene (6 million YBP) in south-western USA and Mexico. By the Pliocene (5 million YBP), the larger Canis lepophagus appeared in the same region and by the Early Pleistocene (1 million YBP) Canis latrans (the Coyote) was in existence. They proposed that the progression from Eucyon davisi to C lepophagus to the Coyote was linear evolution.[5]:p58 Additionally, C. edwardii, C. latrans and C. aureus form together a small clade and because C. edwardii appeared earliest spanning the mid-Blancan (late Pliocene) to the close of the Irvingtonian (late Pleistocene) it is proposed as the ancestor.[19]:p175,180
Nowak and Tedford also believed that it was possible for C. lupus to have been derived from a Miocene or Pliocene canid line that preceded and was separate from C. lepophagus.[7][20] Wang and Tedford proposed that assuming the geological attribution of the material was correct then the earliest identifiable C. lupus remains date 800,000 YBP[19]:5 (Middle Pleistocene) with wolves similar to the living species[19]:p150 occurring in both the Olyorian fauna (Early to Middle Pleistocene of Siberia) and in the Cripple Creek Sump fauna (Alaska), which points to an origin of these wolves in Beringia.[19]:p181 Based on morphology from China, the Pliocene wolf C. chihliensis may have been the ancestor for both C. armbrusteri and C. lupus before their migration into North America.[5]:p148[19]:p181 C. chihliensis appears to be more primitive and smaller than C. lupus, and measurements of its skull and teeth are similar to C. lupus but those of its postcranial elements are smaller.[49] C. amrusteri appeared in North America in the Middle Pleistocene and is a wolf-like form larger than any Canis at that time.[7] At the end of the most recent glacial retreat 30,000 YBP, warming melted the glacial barriers across northern Canada allowing arctic mammals to extend their range into mid-latitude North America, including elk, caribou, bison, and the gray wolf.[5]:p61
In Eurasia during the Middle Pleistocene, C. falconeri gave rise to the hypercarnivore genus Xenocyon, which then gave rise to genus Cuon (the dhole) and genus Lycaon (the African hunting dog).[5]:p105,149 Just before the appearance of C. dirus, North America was invaded by genus Xenocyon that was as large as C. dirus and more hypercarnivorous. The fossil record shows them as rare and it is assumed that they could not compete with the newly derived C. dirus.[5]:p60 The large wolf C. antonii from late Pliocene to early Pleistocene China was assessed as being a variation within C. chihliensis,[19]:p197 and the large wolf C. falconeri occurred abruptly in Europe in the Early Pleistocene, perhaps representing a westward extension of C. antonii.[19]:p181
Canis variabilis
In 2012, a study of the wolf-like Canis species of ancient China under the direction of Xiaoming Wang found that these were all quite close to C. lupus in both dental and post-cranial dimensions except for C. variabilis, which was "very strange" compared to other Canis in China as it had much smaller cranio-dental dimensions than earlier and later species.[49] The study concluded that "It is very likely that this species is the ancestor of the domestic dog Canis familiaris, a hypothesis that has been proposed by previous authors."[50][51][52][53][54]
Canis c.f. familiaris (Paleolithic "dog")
The Paleolithic dog was a Late Pleistocene canine. They were directly associated with human hunting camps in Europe over 30,000 (YBP) and it is proposed that they were domesticated. They are also proposed to be either a proto-dog and the ancestor of the domestic dog or a type of wolf unknown to science.[55]
Canis lupus familiaris (domestic dog)

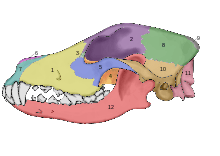
In 2002, a study was undertaken into the fossil skulls of two large canids that had been found buried within meters of the doorway of what was once a mammoth-bone hut at the Eliseevichi-I Upper Paleolithic site in the Bryansk Region on the Russian Plain, and using an accepted morphologically based definition of domestication declared them to be "Ice Age dogs". The carbon dating gave a calendar-year age estimate that ranged between 16,945-13,905 YBP.[56] In 2013, a study looked at one of these skulls and its mitochondrial DNA sequence was identified as Canis lupus familiaris.[57]
In 2015, a zooarchaeologist stated that "In terms of phenotypes, dogs and wolves are fundamentally different animals."[58]
In 1986, a study of skull morphology found that the domestic dog is morphologically distinct from all other canids except the wolf-like canids. "The difference in size and proportion between some breeds are as great as those between any wild genera, but all dogs are clearly members of the same species."[59] In 2010, a study of dog skull shape compared to extant carnivorans proposed that "The greatest shape distances between dog breeds clearly surpass the maximum divergence between species in the Carnivora. Moreover, domestic dogs occupy a range of novel shapes outside the domain of wild carnivorans."[60]
The domestic dog compared to the wolf shows the greatest variation in the size and shape of the skull (Evans 1979) that range from 7 to 28 cm in length (McGreevy 2004). Wolves are dolichocephalic (long skulled) but not as extreme as some breeds of such as greyhounds and Russian wolfhounds (McGreevy 2004). Canine brachycephaly (short-skulledness) is found only in domestic dogs and is related to paedomorphosis (Goodwin 1997). Puppies are born with short snouts, with the longer skull of dolichocephalic dogs emerging in later development (Coppinger 1995). Other differences in head shape between brachycephalic and dolichocephalic dogs include changes in the craniofacial angle (angle between the basilar axis and hard palate) (Regodón 1993), morphology of the temporomandibular joint (Dickie 2001), and radiographic anatomy of the cribriform plate (Schwarz 2000).[61]
Nowak indicated that orbital angle of the eye socket is an important characteristic defining the difference between the dog and the wolf, with the wolf having the lower angle. Nowak compared the orbital angles of four North American canines (including the Indian dog) and produced the following values in degrees: coyote-42.8, wolf-42.8, dog-52.9 dire wolf-53.1. The orbital angle of the eye socket was clearly larger in the dog than in the coyote and the wolf; why it was almost the same as that of the dire wolf was not commented on.[7]
Many authors have concluded that compared to the adult extant wolf, the adult domestic dog has a relatively reduced rostrum (front part of the skull), an elevated frontal bone, a wider palate, a broader cranium, and smaller teeth (Hildebrand1954; Clutton-Brock, Corbet & Hills 1976; Olsen 1985; Wayne 1986; Hemmer 1990; Morey 1990). Other authors have disagreed and have stated that these traits can overlap and vary within the two (Crockford 1999; Harrison 1973). Wolf cubs have similar relative skull proportions as adult dogs and this was proposed as evidence that the domestic dog is a neotenic wolf. This was proposed to be due to either human selection for juvenile appearance or due to a pleiotropic effect as a result of selection for juvenile behavior (Clutton-Brock 1977; Belyaev 1979; Wayne 1986; Coppinger and Schneider 1995). Wayne (1986) concluded that his dog samples did not have significant relative shortening of the rostrum compared to wolves, calling this identification feature into question.[50] A 2004 study that used 310 wolf skulls and over 700 dog skulls representing 100 breeds concluded that the evolution of dog skulls can generally not be described by heterochronic processes such as neoteny although some pedomorphic dog breeds have skulls that resemble the skulls of juvenile wolves.[62] "Dogs are not paedomorphic wolves."[63]
Compared to the wolf, dog dentition is relatively less robust (Olsen 1985; Hemmer 1990), which is proposed to be due to the relaxation of natural selection when wolves became commensal scavengers, or to artificial selection (Olsen 1985; Clutton-Brock 1995). However, Kieser and Groeneveld (1992) compared the mandibulo-dental measurements of jackals (C. adustus, C. mesomelas) and Cape foxes (Vulpes chama) to equivalent-sized dogs and found that the canines of these other canids tended to be slightly smaller and their second molars larger compared to dogs, otherwise the proportions were essentially the same in all species. They concluded that "...the teeth of canids appear to have evolved in concert with one another and relatively independently of differences in dimorphism, size or functional demands". This calls into question the assumption that dog teeth are relatively small due to recent selection, suggesting that dog dentition is plesiomorphic from an ancestor that was smaller than the wolf.[50]
The reduced body size of the early dog compared to a wolf is thought due to niche selection (Olsen 1985; Morey 1992; Coppinger & Coppinger 2001). Morey (1992:199) states that "Results...are consistent with a hypothesis that early domestic dogs are evolutionary paedomorphs, products of strong selection for ontogenetically channeled size reduction and alterations of reproductive timing associated with the new domestic way of life."[50] However, in an domestication experiment the domesticated foxes remained the same size as unselected foxes (Trutt 1999:167).[58]
Wayne (1986) concluded that the dog is closer in skull morphology to C. latrans, C. aureus, C. adustus, C. mesomelas, Cuon alpinus and Lycaon pictus than to the wolf. Dahr (1942) concluded that the shape of the dog brain case is closer to that of the coyote than to that of the wolf. Manwell and Baker (1983) reviewed Dahr's work with the addition of dental data for canids and concluded that the dog ancestor was probably within the range of 13.6–20.5 kg, which is smaller than the range 27–54 kg for extant wolves (Mech 1970) and is comparable with the Dingo.[50]
The auditory bulla of the dog is relatively smaller and flatter than that of the wolf (Harrison 1973; Clutton-Brock, Corbet & Hill 1976; Nowak 1979; Olsen 1985; Wayne 1986), which is proposed to be due to relaxed selection under domestication as the dog no longer required the acute hearing of the wolf. However, bulla shape has been shown to facilitate increased sensitivity to specific frequencies but shape and size may not be correlated with acuity (Ewer 1973). Therefore, the observed difference could be that the dog bulla has retained its ancestral shape.[50]
The ventral edge of the dog's horizontal ramus of the mandible has a convex curve that does not exist in the wolf (Olsen 1985; Clutton-Brock 1995), and no discussion of this difference could be found in the literature. However, Biknevicius and Van Valkenburgh (1997) noticed that the horizontal ramus of bone-processing predators is thicker dorso-ventrally at the point caudal to the site of bone processing. This thickening may have been a function for niche adaptation by the dog's ancestor.[50]
A description of the superficial brain morphology of jackals (C. mesomelas, C. aureus), coyotes (C. latrans), wolves (C. lupus, C. rufus), and dogs indicated that the cerebellum of the dog closely approximates that of the coyote, which is closely aligned with the jackals, and that the wolves show numerous brain traits distinct from the other species (Atkins and Dillon 1971). Wolves also have serological and biochemical traits distinct from dogs (Leone and Wiens 1956; Lauer, Kuyt & Baker 1969).[50]
During the Last Glacial Maximum, there was greater wolf genetic diversity than there is today,[64][57] and within the Pleistocene gray wolf population the variations between local environments would have encouraged a range of wolf ecotypes that were genetically, morphologically and ecologically distinct from one another.[65] One author has proposed that the most likely explanation for the different morphological characteristics of the dog compared to the wolf is that the dog's ancestor was adapted to a different niche than the wolf.[50]
Genetic record
DNA sequences
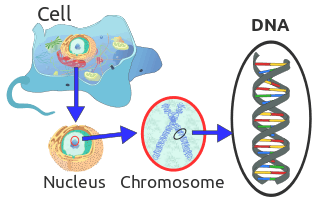
The mitochondria within each cell contain many copies of a small circular DNA genome and in mammals it is 16,000–18,000 base pairs in length. A cell contains hundreds or thousands of mitochondria and therefore the genes contained within those mitochondria are more abundant than the genes that occur in the nucleus of the cell.[66][67] The abundance of mitochondrial DNA (mDNA) is useful for the genetic analysis of ancient remains where the DNA has degraded.[67][68]
Mitochondrial DNA sequences have a higher mutation rate than the mutation rate of nuclear genes and for mammals this rate is 5–10 times faster.[67][69][70] The mitochondrial protein-coding genes evolve much faster and are powerful markers for inferring evolution history at category levels such as families, genera, and species. However, they have evolved at a faster rate than other DNA markers and there is a timing difference in its molecular clock that needs to be validated against other sources. The taxonomic status of uncertain species is better resolved through using nuclear DNA, which is more suitable for analyzing the recent history.[71] In most cases, mDNA is inherited from the maternal ancestor.[67][72] Therefore, phylogenetic analysis of mDNA sequences within species provides a history of maternal lineages that can be represented as a phylogenetic tree.[67][73][74]
| From specimen to phylogenetic tree | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
| Phylogenetic tree | ||||||||||||||||||||||||
| ||||||||||||||||||||||||
| Phylogenetic relationship between four canids.[76][77] |
The mutations that are different in these 4 sequences have been numbered and bolded. These mutations can then be used to build a phylogenetic tree for the four canids. In this example, the dog and gray wolf differ by two substitutions (highlighted in red), and each of them differs from the coyote by four substitutions.[67]
1 2 3 4 5 67
Golden Jackal A-G-C-T-G-T-C-GA-T-TC-CA
Coyote A-G-C-T-A-T-C-GA-A-TC-GA
Wolf T-G-C-T-A-T-G-GA-T-TC-CT
Dog T-G-G-T-A-T-G-GA-T-TC-CA
The mDNA sequences of the dog and wolf differ by only 0–12 substitutions within 261 base-pairs, whereas dogs always differed from coyotes and jackals by at least 20 substitutions.[67][78] This finding implies that the dog derived from the wolf and that there has been repeated back-crossing,[78] or that the dog may have descended from a now extinct species of canid whose closest living relative is the modern wolf.[79]
Timing issue
DNA studies may give unresolvable results due to the specimens selected, the technology used, and the assumptions made by the researchers.[80] Any one from a panel of genetic markers can be chosen for use in a study, and the techniques used to extract, locate and compare sequences can be applied using advances in technology to observe longer lengths of base pairs that give better phylogenetic resolution.[81] Phylogenetic trees based on different genetic markers have given conflicting results regarding the relationship between the wolf, dog and coyote. One study based on SNPs[82] and another based on nuclear gene sequences[83] showed dogs clustering with coyotes and separate from wolves. Another study based on SNP genotypes showed wolves clustering with coyotes and separate from dogs.[84] Other studies based on a number of markers show the more widely accepted result of wolves clustering with dogs separate from coyotes.[78][85] These results indicate that caution is needed when interpreting the results of genetic markers.[82]
There are two key assumptions that are made for dating the divergence time for species: the generation time and the genetic mutation rate per generation. The time between generations for wolves is assumed to be three years based on the extant gray wolf, and two years for the dog based on the extant dog.[76] One recent major study assumed a generation time of 2 years for the dog for as far back as 10,000 years ago, and then assumed a generation time of 3 years (the same as the wolf) before that to calculate a proposed divergence time between the two.[64]
DNA studies are conducted but with "the mutation rate as the dominant source of uncertainty."[64] In 2005, Lindblad-Toh sequenced the first draft genome of the extant dog, and calculated a proposed mutation rate of 1x10−8 mutations per generation.[76] In 2015, Skoglund was able to sequence the first draft genome of the 35,000 YBP Taimyr wolf and used its radio-carbon date to validate a proposed genetic mutation rate of 0.4x10−8 mutations per generation.[86] The difference is a timing factor of 2.5, however another study stated that because only one Pleistocene wolf specimen has so far been sequenced, then the result should be treated with caution, with that study then providing both estimates to calculate the proposed divergence times between the wolf and dog.[87] However, in 2016 the mutation rate of the 4,800 YBP Newgrange dog matched that of the Taimyr wolf.[88]
Wolf-like canids
| The extant wolf-like canids | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Phylogenetic relationships between the extant wolf-like clade of canids based on mDNA.[76][77] |
- Further information: Canid phylogeny, and Canid hybrid
The wolf-like canids are a group of large carnivores that are genetically closely related because their chromosomes number 78. The group includes genus Canis, Cuon and Lycaon. The members are the dog (C. lupus familiaris), gray wolf (C. lupus), coyote (C. latrans), golden jackal (C. aureus), Ethiopian wolf (C. simensis), black-backed jackal (C. mesomelas), side-striped jackal (C. adustus), dhole (Cuon alpinus), and African wild dog (Lycaon pictus).[89] Newly proposed members include the red wolf (Canis rufus), eastern wolf (Canis lycaon), and African golden wolf (C. anthus). As they possess 78 chromosomes, all members of the genus Canis (coyotes, wolves, jackals) are karyologically indistinguishable from each other, and from the dhole and the African hunting dog.[67]:p279[90] The members of Canis can potentially interbreed[79] and there is evidence that the Ethiopian wolf has hybridized with dogs.[91] According to zoologist Reginald Pocock, a dhole interbred with a golden jackal.[92] The African hunting dog is large, highly mobile, known to disperse over large distances and are rare throughout much of their geographical range,[93] making opportunities for hybridization difficult. A study of the maternal mitochondrial DNA of the black-backed jackal could find no evidence of genotypes from the most likely mates – the side-striped jackal nor the golden jackal – indicating that male black-backed jackals had not bred with these.[94] A search of the scientific literature could not find evidence of hybridization for the rare side-striped jackal.
A DNA sequence alignment for the wolf-like canids gave a phylogenetic tree with the gray wolf and dog being the most closely related, followed by a close affiliation with the coyote, golden jackal and Ethiopian wolf, and the dog can hybridize in the wild with these three species. Next closest to this group are the dhole and African wild dog that both have unique meat-slicing teeth, suggesting that this adaptation was later lost by the other members.[76] The two African jackals are shown as the most basal members of this clade, which means that this tree is indicating an African origin for the clade.[76][95] The tree illustrates the genotype-phenotype distinction, where a genotype is an organism's full hereditary information and a phenotype is an organism's actual observed properties, such as morphology, development, or behavior. By phenotype, the dhole (genus Cuon) and the African hunting dog (genus Lycaon) are not classified as members of the genus Canis, but by genotype they are closer to dogs, wolves and coyotes than are the two genus Canis jackals - the Side-striped jackal (C. adustus) and the Black-backed jackal (C. mesomelas).
In 2014, a whole-genome DNA study indicated that the golden jackal ancestral lineage had diverged from the wolf/coyote ancestral lineage 400,000 years ago, which is considerably more recent than previous estimates of 1.9 million years based on mitochondrial DNA, but with "the mutation rate as the dominant source of uncertainty."[64] Using an alternate mutation rate per generation (Skoglund 2015),[86] the divergence time becomes 955,000-1 million YBP.
In 2015, a study of mitochondrial genome sequences and nuclear genome sequences of African and Eurasian canids indicated that extant wolf-like canids had colonized Africa from Eurasia at least 5 times throughout the Pliocene and Pleistocene, which is consistent with fossil evidence suggesting that much of the African canid diversity resulted from the immigration of Eurasian ancestors, likely coincident with Plio-Pleistocene climatic oscillations between arid and humid conditions. When comparing the African and Eurasian golden jackals, the study concluded that the African specimens represented a distinct monophyletic lineage that should be recognized as a separate species, Canis anthus (African golden wolf). According to a phylogeny derived from nuclear sequences, the Eurasian golden jackal (Canis aureus) diverged from the wolf/coyote lineage 1.9 million years ago but the African golden wolf separated 1.3 million years ago. Mitochondrial genome sequences indicated the Ethiopian wolf diverged from the wolf/coyote lineage slightly prior to that.[77]
Two wolf haplogroups
| mDNA Haplogroups | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Divergence times and phylogenetic mDNA relationships. Haplogroup 2 basal to and 5 mutations from Haplogroup 1,[97] dog is closer to the ancient wolves of Western Europe than to modern wolves.[57] |
A haplotype (haploid genotype) is a group of genes in an organism that are inherited together from a single parent.[99][100] A haplogroup is a group of similar haplotypes that share a common ancestor with a single-nucleotide polymorphism mutation.[101][102] Mitochondrial DNA passes along a maternal lineage that can date back thousands of years.[101]
In 2010, a study compared a 230 base pair sequence of the mitochondrial control region from 24 ancient wolf specimens from western Europe dated between 44,000–1,200 YBP with those of modern gray wolves. Most of the sequences could be represented on a phylogenetic tree. However, the haplotypes of the Himalayan wolf and the Indian gray wolf could not because they were 8 mutations apart from the other wolves,[97] indicating distinct lineages which had previously been found in other studies.[97][96][103] The study found that there were 75 different gray wolf mDNA haplotypes that include 23 in Europe, 30 in Asia, 18 in North America, 3 in both Europe and Asia, and 1 in both Europe and North America.[97] These haplotypes formed two haplogroups that were separated from each other by 5 mutations. Haplogroup 1 formed a monophyletic clade (indicating that they all carried the same mutation inherited from a single female ancestor). All other haplotypes were basal in the tree, and these formed 2–3 smaller clades that were assigned to haplogroup 2 that was not monophyletic.[97][104]
Haplogroups 1 and 2 could be found spread across Eurasia but only haplogroup 1 could be found in North America. The ancient wolf samples from western Europe all belonged to haplogroup 2, indicating haplogroup 2 predominance in this region for over 40,000 years before and after the Last Glacial Maximum. A comparison of current and past frequencies indicated that in Europe haplogroup 2 became outnumbered by haplogroup 1 over the past several thousand years but in North America haplogroup 2 became extinct and was replaced by haplogroup 1 after the Last Glacial Maximum.[97][104] Access into North America was available between 20,000–11,000 years ago, after the Wisconsin glaciation had retreated but before the Bering land bridge became inundated by the sea.[105] Therefore, haplogroup 1 was able to enter into North America during this period.
Stable isotope analysis conducted on the bone of a specimen allows researchers to form conclusions about the diet, and therefore the ecology, of extinct wolf populations. This analysis suggests that the Pleistocene wolves from haplogroup 2 found in Beringia and Belgium preyed mainly on Pleistocene megafauna,[55][97][106] which became rare at the beginning of the Holocene 12,000 years ago.[97][107] One study found the Beringian wolf to be basal to all other gray wolves except for the extant Indian gray wolf and the extant Himalayan wolf.[106] The Pleistocene Eurasian wolves have been found to be morphologically and genetically comparable to the Pleistocene eastern-Beringian wolves,[108] with some of the ancient European and Beringian wolves sharing a common haplotype (a17),[97][106] which makes ecological similarity likely.[97] Two ancient wolves from the Ukraine dated around 30,000 YBP and the 33,000 YBP "Altai dog" had the same sequence as six Beringian wolves, and another from the Czech Republic dated 44,000 YBP had the same sequence as two Beringian wolves.[106]
It has been proposed that the Pleistocene wolves across northern Eurasia and northern North America represented a continuous and almost panmictic population that was genetically and probably also ecologically distinct from the wolves living in this area today.[97][109] The specialized Pleistocene wolves did not contribute to the genetic diversity of modern wolves, and the modern wolf populations across the Holarctic are likely to be the descendants of wolves from populations that came from more southern refuges.[109] Extant haplogroup 2 wolves can be found in Italy, the Balkans and the Carpathian Mountains but rare elsewhere in Europe. In Asia, only four haplotypes have been identified as belonging to this haplogroup, and two of them occur in the Middle East.[110] Haplogroup 2 did not become extinct in Europe, and if before the Last Glacial Maximum haplogroup 2 was exclusively associated with the wolf ecomorph specialized in preying on megafauna, it would mean that in Europe it was capable of adapting to changing prey.[97]
In 2013, a mitochondrial DNA sequencing of ancient wolf-like canids revealed another separate lineage of 3 haplotypes (forming a haplogroup) that was found in 3 Late Pleistocene specimens from Belgium; however, it has not been detected in extant wolves.[57][110] One of these was the "Goyet dog".[57]
Dissenting view
In 2016, a study was undertaken due to concerns that previous mDNA studies may have been conducted with insufficient genetic resolution or limited geographical coverage and had not included sufficient specimens from Russia, China, and the Middle East. The study compared a 582 base pair sequence of the mitochondrial control region which gave twice the phylogenetic resolution of the 2010 study.[97] The study compared the sequences of both modern wolves and ancient wolf specimens, including specimens from the remote areas of North America, Russia and China. The study included the Taimyr wolves, the Goyet "dog", the Altai "dog", Beringian wolves, and other ancient specimens.[111]
The study found 114 different wolf haplotypes among 314 sequences, with the new haplotypes being found in Siberia and China. The phylogenetic tree resolved into 19 clades that included both modern and ancient wolves, which showed that the most basal clades included the Indian gray wolf and the Himalayan wolf, with a subclade of wolves from China and Mongolia falling within the Himalayan wolf clade. The two most basal North American haplotypes included the Mexican wolf and the Vancouver Island wolf. In Europe, the two most genetically distinct haplotypes form the Italian wolf and separately the Iberian wolf. The Greenland wolves all belonged to one haplotype that had been previously found among North American wolves and which indicates their origin from North America. The Eastern wolf was confirmed as a coyote/wolf hybrid. Wolves found in the regions of the Chukotka Peninsula, the North Korean border, Amur Oblast and Khakassia showed the greatest genetic diversity and with close links to all other wolves found across the holarctic. One ancient haplotype that had been found in Alaska (Eastern Beringia 28,000 YBP) and Russia (Medvezya Cave 18,000 YBP) was shared with some modern wolves found in China and Mongolia.[111]
The previous finding of two wolf haplogroups[97] was not clearly delineated in this study but it agreed that the genetic diversity of past wolves has been lost at the beginning of the Holocene in Alaska, Siberia, and Europe with limited overlap with modern wolves. For the ancient wolves of North America, instead of an extinction/replacement model suggested by a previous study,[106] this study found substantial evidence of a population bottleneck in North America in which the ancient wolf diversity was almost lost around the beginning of the Holocene (no further elaboration in the study). In Eurasia, the loss of ancient lineages could not be simply explained and appears to have been slow across time with the reasons unclear.[111]
Into America and Japan
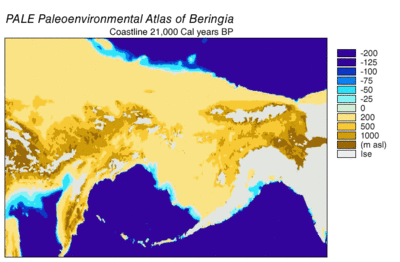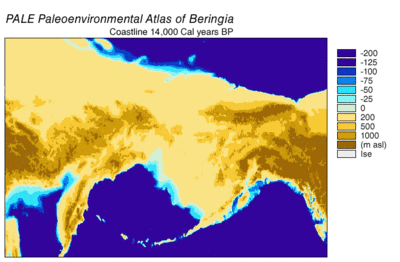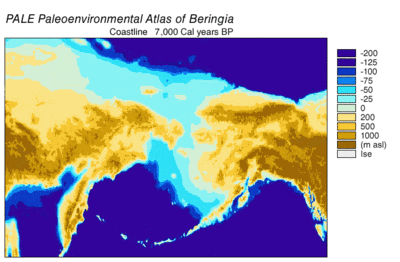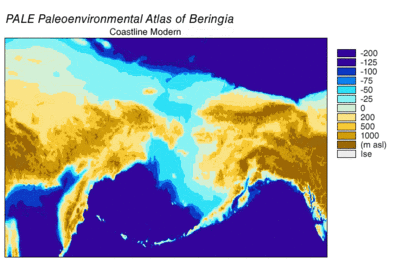
In 2016, a study built on the work of another major study[57] and analyzed the sequences of 12 genes that are located on the heavy strand of the mitochondrial genome of extinct and modern C. lupus. The study excluded the sequences of the divergent Himalayan wolf and the Indian gray wolf. The ancient specimens were radiocarbon dated and stratagraphically dated, and together with the sequences generated a time-based phylogenetic tree. From the tree, the study was able to infer the most recent common ancestor for all other C. lupus specimens - modern and extinct - was 80,000 YBP and this date concured with the earlier study.[57][112] The study could find no evidence of a population bottleneck for wolves until a few thousand years ago.[112]
The phylogenetic tree showed the polyphyly of American wolves, the Mexican wolf was divergent from other North American wolves, and these other North American wolves formed two closely related clades. A scenario consistent with the phylogenetic, ice sheet and sea-level data was that during the Ice Age when sea levels were at their lowest, there was a single wave of wolf colonization into North America starting with the opening of the Bering land bridge 70,000 YBP and closing during the Late Glacial Maximum of the Yukon corridor that ran through the division between the Laurentide Ice Sheet and the Cordilleran Ice Sheet 23,000 YBP. Mexican wolves were part of the single wave and either diverged from the other wolves before entering North America or once in North America due to the change in its environment.
As wolves had been in the fossil record of North America but modern wolves could trace their ancestry back only 80,000 years, the wolf haplotypes that were already in North America were replaced by these invaders, either through competitive displacement or through admixture.[112] The replacement in North America of a basal population of wolves by a more recent one supported the findings of earlier studies.[97][104][106][112] There possibly existed a panmictic wolf population with gene flow spanning Eurasia and North America until the closing of the ice sheets.[97][109][112] Once the sheets closed, the southern wolves were isolated and north of the sheets only the Beringian wolf existed. The land bridge became inundated by the sea 10,000 YBP, the sheets receded 12,000–6,000 YBP, the Beringian wolf went extinct and the southern wolves expanded to recolonize the rest of North America. All North American wolves are descended from those that were once isolated south of the ice sheets. However, much of their diversity was later lost during the twentieth century.[112]
Studies using mitochondrial DNA have indicated that the wolves of coastal south-east Alaska are genetically distinct from inland gray wolves, reflecting a pattern also observed in other taxa. They show a phylogenetic relationship with extirpated wolves from the south (Oklahoma), indicating that these wolves are the last remains of a once widespread group that has been largely extirpated during the last century, and that the wolves of northern North America had originally expanded from southern refuges below the Wisconsin glaciation after the ice had melted at the end of the Last Glacial Maximum.[113][114][115] A whole-genome DNA study indicated that all North American wolves were monophyletic and therefore are the descendants of a common ancestor.[116]
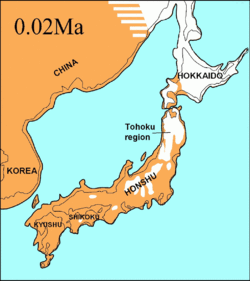
During the same period, the Soya Strait between Hokkaido and Sakhalin Island was dry for 75,000 years and it was proposed that the extinct Ezo wolf (C. l. hattai) arrived on Hokkaido from Sakhalin.[98][112][117] However, the sequences indicated that it arrived in Hokkaido less than 10,000 YBP. The Ezo wolf was closely related to one of the North American clades,[98][112][118] but different to the more southerly Japanese wolf (C. l. hodophilax) that was basal to modern wolves.[98][112] The Japanese wolf inhabited Kyushu, Shikoku, and Honshu islands[119][120] but not Hokkaido Island.[120] This indicates that its ancestor may have migrated from the Asian continent through the Korean Peninsula into Japan.[98][120] The past sea levels of the Korean Strait together with the timing of the Japanese wolf sequences indicated that it arrived to the southern islands less than 20,000 YBP.[112]
The dog was a very successful invader of North America and had established a widespread ecological niche by the Early–Middle Holocene. There was no overlap in niche between the dog and the wolf in comparison to the dog and other North American canids. By the Late Holocene, the dog's niche area was less in size than researchers had expected to find, indicating that it was limited by biotic factors. These regions include the northeast and northwest of the United States that correlate with the greatest densities of early human occupation, indicating that the dog had "defected" from the wolf niche to the human niche and explains why the dog's niche area was not as large as expected. The separation between dog and wolf may reflect the rapid rate in which domestication occurred,[121] including the possibility of a second domestication event occurring in North America.[122][121] Packs of wolves and hunter-gatherers hunt similar prey in a similar way within a similar group social structure that may have facilitated wolf domestication.[123][51]
Divergence with the coyote
In 2016, a whole-genome DNA study proposed, based on the assumptions made, that all of the North American wolves and coyotes diverged from a common ancestor less than 6,000–117,000 years ago. The study also indicated that all North America wolves have a significant amount of coyote ancestry and all coyotes some degree of wolf ancestry, and that the red wolf and eastern wolf are highly admixed with different proportions of gray wolf and coyote ancestry. One test indicated a wolf/coyote divergence time of 51,000 years before present that matched other studies indicating that the extant wolf came into being around this time. Another test indicated that the red wolf diverged from the coyote between 55,000-117,000 years before present and the Great Lakes region wolf 32,000 years before present. Other tests and modelling showed various divergence ranges and the conclusion was a range of less than 6,000 and 117,000 years before present.[116][124] This finding conflicts with the fossil record that indicates a coyote-like specimen dated to 1 million years before present.[5]
Domestic dog
The domestic dog (Canis lupus familiaris) is the most widely abundant large carnivore.[57][87][125] Over the past million years, numerous wolf-like forms existed but their turnover has been high, and modern wolves are not the lineal ancestors of dogs.[57][64][87][126] Although research had suggested that dogs and wolves were genetically very close relatives,[78][79][89] later phylogenetic analysis strongly supported the hypothesis that dogs and wolves are reciprocally monophylic taxa that form two sister clades,[78][64][127] suggesting that none of the modern wolf populations are related to the wolves that were first domesticated.[64][127] Recent mitochondrial DNA analyses of ancient and modern gray wolf specimens supports a pattern of population reduction and turnover.[57][97][106]
In 2016, a study investigated for the first time the population subdivisions, demography, and the relationships of gray wolves based on their whole-genome sequences. The study indicated that the dog was a divergent subspecies of the gray wolf and was derived from a now-extinct ghost population of Late Pleistocene wolves,[57][64][87] and the dog and the dingo are not separate species.[87] The genome-wide phylogenetic tree indicated a genetic divergence between New World and Old World wolves, which was then followed by a divergence between the dog and Old World wolves 27,000YBP[86][87] - 29,000 YBP.[87] The dog forms a sister taxon with Eurasian gray wolves but not North American wolves. The dog had considerable pre-ancestry after its divergence from the Old World wolves before it separated into distinct lineages that are nearly as distinct from one another as they are from wolves.[87] The study suggested that previous datings based on the divergence between wolves and coyotes of one million years ago using fossils of what appeared to be coyote-like specimens may not reflect the ancestry of the modern forms.[77][64][86][87]
| Gray wolf divergence and timing | |||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||
| Whole-genome phylogenetic tree - extant gray wolf populations,[87] with divergence times calculated using an assumed mutation rate of Lindblad-Toh (1x10−8)[76] or [Skoglund] (0.4x10−8).[86] |
The study indicated that the Mexican wolf was also a divergent form of gray wolf, suggesting that may have been part of an early invasion into North America.[87][126] The Tibetan wolf was found to be the most highly divergent of the Old World wolves, had suffered a historical population bottleneck and had only recently recolonized the Tibetan Plateau. Glaciation may have caused its habitat loss, genetic isolation then local adaption.[87]
The study indicated that there has been extensive genetic admixture between domestic dogs and wolves, with up to 25% of the genome of Old World wolves showing signs of dog ancestry, possibly as the result of gene flow from dogs into wolves that were ancestral to all modern wolves. There was evidence of significant gene flow between the European wolves plus the Israeli wolf with the basenji and boxer, which suggests admixture between the lineages ancestral to these breeds and wolf populations.[64][87] For the lowland Asian wolves: the Central Russian and East Russian wolves and all of the lowland Chinese wolves had significant gene flow with the Chinese indigenous dogs, the Tibetan Mastiff and the dingo. For the highland Asian wolves: The Tibetan wolves did not show significant admixture with dogs; however, the Qinghai wolves had gene flow with the dingo and one of them had gene flow with the Chinese dogs. The New World wolves did not show any gene flow with the boxer, dingo or Chinese indigenous dogs but there was indication of gene flow between the Mexican wolf and the African basenji.[87] All species within the Canis genus, the wolf-like canids, are phylogenetically closely related with 78 chromosomes and can potentially interbreed.[79] There was indication of gene flow into the golden jackal from the population ancestral to all wolves and dogs (11.3%–13.6%) and much lower rates (up to 2.8%) from extant wolf populations.[64][87]
The data indicated that all wolves shared similar population trajectories, followed by population decline that coincided with the expansion of modern humans worldwide and their technology for capturing large game.[87][128] Late Pleistocene carnivores would have been social living in large prides, clans and packs in order to hunt the larger game available at that time, and these larger groups would have been more conspicuous targets for human persecutors.[128] Large dogs accompanying the humans may have accelerated the rate of decline of carnivores that competed for game,[87][129] therefore humans expanded across Eurasia, encountered wolves, domesticated some and possibly caused the decline of others.[87]
The study concluded that admixture had confounded the ability to make inferences about the place of dog domestication. Past studies based on SNPs, genome-wide similarities with Chinese wolves, and lower linkage disequilibrium might reflect regional admixture between dogs with wolves and gene flow between dog populations, with divergent dog breeds possibly maintaining more wolf ancestry in their genome. The study proposed that analysis of ancient DNA might be a better approach.[87]
In the same year, a study found that there were only 11 fixed genes that showed variation between wolves and dogs. These genes are thought to affect tameness and emotional processing ability.[130] Another study provided a listing of all of the gray wolf and dog mDNA haplotypes combined in the one phylogenetic tree.[131]
Taimyr wolf

In May 2015, a study was conducted on a partial rib-bone of a wolf specimen (named Taimyr-1) found near the Bolshaya Balakhnaya River in the Taimyr Peninsula of Arctic North Asia, that was AMS radiocarbon dated to 34,900 YBP. The sample provided the first draft of the nuclear genome of a Pleistocene carnivore, and the sequence was deposited in the European Nucleotide Archive and classified as Canis lupus.[86]
Using the Taimyr-1 specimen's radiocarbon date, its genome sequence and that of a modern wolf, a direct estimate of the genome-wide mutation rate in dogs/wolves could be made to calculate the time of divergence. The data indicated that the previously unknown Taimyr-1 lineage was a wolf population separate to modern wolves and dogs and indicated that the Taimyr-1 genotype, gray wolves and dogs diverged from a now-extinct common ancestor[86][65][132] before the peak of the Last Glacial Maximum 27,000-40,000 years ago. The separation of the dog and wolf did not have to coincide with selective breeding by humans.[86][133] Such an early divergence is consistent with several paleontological reports of dog-like canids dated up to 36,000 YBP, as well as evidence that domesticated dogs most likely accompanied early colonizers into the Americas.[86]
Comparison to the gray wolf lineage indicated that Taimyr-1 was basal to gray wolves from the Middle East, China, Europe and North America but shared a substantial amount of history with the present-day gray wolves after their divergence from the coyote. This implies that the ancestry of the majority of gray wolf populations today stems from an ancestral population that lived less than 35,000 years ago but before the inundation of the Bering Land Bridge with the subsequent isolation of Eurasian and North American wolves.[86]
A comparison of the ancestry of the Taimyr-1 lineage to the dog lineage indicated that some modern dog breeds have a closer association with either the gray wolf or Taimyr-1 due to admixture. The Saarloos wolfdog showed more association with the gray wolf, which is in agreement with the documented historical crossbreeding with gray wolves in this breed. Taimyr-1 shared more alleles (gene expressions) with those breeds that are associated with high latitudes - the Siberian husky and Greenland dog[86][132] that are also associated with arctic human populations, and to a lesser extent the Shar Pei and Finnish spitz. An admixture graph of the Greenland dog indicates a best-fit of 3.5% shared material, although an ancestry proportion ranging between 1.4% and 27.3% is consistent with the data. This indicates admixture between the Taimyr-1 population and the ancestral dog population of these four high-latitude breeds. These results can be explained either by a very early presence of dogs in northern Eurasia or by the genetic legacy of Taimyr-1 being preserved in northern wolf populations until the arrival of dogs at high latitudes. This introgression could have provided early dogs living in high latitudes with phenotypic variation beneficial for adaption to a new and challenging environment. It also indicates that the ancestry of present-day dog breeds descends from more than one region.[86]
An attempt to explore admixture between Taimyr-1 and gray wolves produced unreliable results.[86]
As the Taimyr wolf had contributed to the genetic makeup of the Arctic breeds, a later study suggested that descendants of the Taimyr wolf survived until dogs were domesticated in Europe and arrived at high latitudes where they mixed with local wolves, and these both contributed to the modern Arctic breeds. Based on the most widely accepted oldest zooarchaeological dog remains, domestic dogs most likely arrived at high latitudes within the last 15,000 years. The mutation rates calibrated from both the Taimyr wolf and the Newgrange dog genomes suggest that modern wolf and dog populations diverged from a common ancestor between 20,000 and 60,000 YBP. This indicates that either dogs were domesticated much earlier than their first appearance in the archaeological record, or they arrived in the Arctic early, or both.[134]
The finding of a second wolf specimen from the same area (Taimry-2) and dated to 42,000 YBP has also been reported but only yielded mitochondrial DNA.[135]
Canis variabilis
In 2015, a study looked at the mitochondrial control region sequences of 13 ancient canid remains and one modern wolf from five sites across Arctic north-east Siberia. The fourteen canids revealed nine mitochondrial haplotypes, three of which were on record and the others not reported before. The phylogentic tree generated from the sequences showed that four of the Siberian canids dated 28,000 YBP and one Canis c.f. variabilis dated 360,000 YBP were highly divergent. The haplotype designated as S805 (28,000 YBP) from the Yana River was one mutation away from another haplotype S902 (8,000 YBP) that represents Clade A of the modern wolf and domestic dog lineages. Closely related to this haplotype was one that was found in the recently-extinct Japanese wolf. Several ancient haplotypes were oriented around S805, including Canis c.f. variabilis (360,000 YBP), Belgium (36,000 YBP - the "Goyet dog"), Belgium (30,000 YBP), and Konsteki, Russia (22,000 YBP). Given the position of the S805 haplotype on the phylogenetic tree, it may potentially represent a direct link from the progenitor (including Canis c.f. variabilis) to the domestic dog and modern wolf lineages. The gray wolf is thought to be ancestral to the domestic dog, however its relationship to C. variabilis, and the genetic contribution of C. variabilis to the dog, is the subject of debate.[136]
The Zhokhov Island (8,700 YBP) and Aachim (1,700 YBP) canid haplotypes fell within the domestic dog clade, cluster with S805, and also share their haplotypes with - or are one mutation away from - the Tibetan wolf (C. l. chanco) and the recently-extinct Japanese wolf (C. l. hodophilax). This may indicate that these canids retained the genetic signature of admixture with regional wolf populations. Another haplotype designated as S504 (47,000 YBP) from Duvanny Yar appeared on the phylogenetic tree as not being connected to wolves (both ancient and modern) yet ancestral to dogs, and may represent a genetic source for regional dogs.[136]
Rise to dominant predator
In 2015, a study looked at the paleoecology of large carnivores across the Mammoth steppe during the Late Pleistocene by using stable isotope analysis of their fossil collagen to reconstruct their diets. Based on testing in Belgium, around 40,000 YBP the Cave hyenas preyed on mammoth, woolly rhinoceros, horses and reindeer, with cave lions taking reindeer and young cave bears. Wolves appear to have been out-competed by cave hyenas and had their diet restricted to chamois, giant deer and red deer. However, after the Last Glacial Maximum around 14,000 YBP, wolves had access to all prey species, the cave lion was restricted to reindeer, and the cave hyena had gone extinct.[137][138][139] The data suggests that the extinction of the cave hyena allowed the wolf to become the dominant predator rather than the cave lion, just before the cave lion's extinction.[139] Another study indicated that the wolf thrived compared to the cave hyena when there was greater snow cover.[140]
Wolf population differences
Gray wolves have a wide, natural distribution across the Holarctic that includes many different habitats, which can vary from the high arctic to dense forests, open steppe and deserts. The genetic differences between different populations of gray wolves is tightly linked to the type of habitat in which they live.[141] Differences in genetic markers among the Scandinavian wolf population has arisen in only just over a decade due to their small population size,[141][142] which indicates that these differences are not dependent on a long time spent in isolation and that larger population patterns can evolve in just a few thousand years.[141] These differences can also include fur color and density, and body size.[141][143][144] The differences can also include behavior, as coastal wolves eat fish[141][143] and tundra wolves migrate.[141][144] These differences have been observed between two wolf populations that are living in close proximity. It has been shown that mountain wolves do not interbreed with nearby coastal wolves, and the Alps of France and Switzerland have been repopulated with wolves from the mountains of nearby Italy[141][145] and from the far away mountains of Croatia[141][146] rather than from the nearer lowlands, which indicates that distance is not the driving force in differences between the two ecomorphs.[141]
Ecological factors including habitat type, climate, prey specialization and predatory competition will greatly influence their genetic population structure and cranio-dental plasticity.[107][65][141][144][147][148][149][150][151] Within the Pleistocene gray wolf population the variations between local environments would have encouraged a range of wolf ecotypes that were genetically, morphologically and ecologically distinct from one another.[65]
Ecotypes
In 2016, two studies compared the sequences of 42,000 single nucleotide polymorphisms in North American gray wolves and found that they formed 6 ecotypes - a genetically and ecologically distinct population separated from other populations by their different type of habitat. These six wolf ecotypes were named West Forest, Boreal Forest, Arctic, High Arctic, Baffin, and British Columbia. The studies found that precipitation and mean diurnal temperature range were the most influential variables on sequence variation.[152][153] These findings were in accord with previous findings that precipitation influenced morphology,[154] and that vegetation[149] and habitat type[143][155] influenced wolf differences. One of these studies found that the variation in 11 key genes had an impact on wolf vision, sense of smell, hearing, coat color, metabolism, and immunity. The study identified 1,040 genes that are potentially under selection due to habitat variation, and therefore that there was evidence of local adaption of the wolf ecotypes at a molecular level. Most notable was the positive selection of genes that influence vision, coat color, metabolism and immunity in the Artic and High Arctic ecotypes, and that the British Columbia ecotype also has a unique set of adaptions.[153] The local adaptation of a wolf ecotype most likely reflects a wolf’s preference to remain in the type of habitat that it was born into.[152]
Pleistocene wolves
The oldest Canis remains found in Europe were from France and dated to 3.1 million YBP,[156] followed by Canis cf. etruscus (where cf. in Latin means confer, uncertain) from Italy dated to 2.2 million YBP.[157] C. lupus first appeared in Italy during Marine Isotope Stage 9[158] (337,000 YBP). In Britain, it was the only canid species present from MIS 7 (243,000 YBP), with the oldest record from Pontnewydd Cave in north Wales.[159] During the Ice Age, Britain was separated from Europe by only the Channel River.
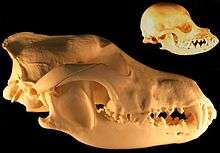
A study of Pleistocene C. lupus in Britain at different time periods found that its abilities to crush, slice meat and eat bone highlighted its cranio-dental plasticity. These responses to dietary changes showed species-wide dietary shifts, and not just local ecomorphs, in response to climatic and ecological variables. The survival of C. lupus during the Pleistocene can be attributed largely to its plastic cranio-dental morphology.[151]


| Time YBP | Variables |
|---|---|
| 243,000 MIS 7 | Paleoenvironment was open grasslands with summer temperatures between 16 °C and 23 °C and winter temperatures between −7 °C and −6 °C dominated by steppe mammoth and horse. Competitors included the lion, brown bear, and rarely the spotted hyena. The wolves of MIS 7 were slightly smaller in body size than MIS 5 wolves and those found in Sweden today. These wolves were out-competed by the larger competitors, leading to a more omnivorous diet with increased crushing ability in an open environment that supported more types of prey and more non-meat foods than the MIS 5 period. They had shallower and narrower jaws than MIS 5 wolves and those found in Sweden today, which indicated that they could take only small to medium-sized prey. They exhibited a lower percentage of tooth breakage comparable with MIS-3 wolves. However, they had the highest percentage of moderately worn teeth.[151] |
| 82,000 MIS 5A | Paleoenvironment was cold, open tundra with summer temperatures between 7 °C and 11 °C and winter temperatures between −10 °C and −30 °C dominated by reindeer and bison. A large form of brown bear was top predator, with no hyena at this time. The wolves of MIS 5 were larger in body size than those found in Sweden today. These wolves suffered from a severe climate, low prey availability and dietary stress leading to a more carnivorous diet, with increased scavenging of frozen carcasses and bone consumption. They developed strong jaws and the highest flesh-slicing ability compared to the other wolves, with shallower jaws than the modern wolf but broader and deeper jaws than MIS 7 and MIS 5 wolves. They exhibited the longest and narrowest upper P4 that suggests improved slicing ability, and longest upper M1 and M2 but with reduced width and therefore reduced crushing ability, indicating a hypercarnivore. They exhibited a higher percentage of tooth breakage and severely worn teeth compared to the other wolves, and may have been using their upper P4 and lower m1 to crush bone rather than their molars, leading to a higher frequency of damage.[151] |
| 57,000 MIS 3 | Paleoenvironment of open grasslands with summer temperature of around 12 °C and winter temperature around −20 °C dominated by woolly mammoth, woolly rhinoceros, horse, and giant deer. Competitors included the lion, brown bear, and the spotted hyena as the top carnivore. The wolves of MIS 3 were smaller in body size than MIS 5 wolves and those found in Sweden today. These wolves were out-competed by the lion and hyena, leading to a more omnivorous diet with increased crushing ability in an open environment that supported more types of prey and more non-meat foods than the MIS 5 period. They had shallower and narrower jaws than MIS 5 wolves and those found in Sweden today, which indicated that they could take only small to medium-sized prey. They exhibited a lower percentage of tooth breakage comparable with MIS-7 wolves with moderate tooth wear.[151] |
| Today (Sweden) | Wolves have been extirpated in Britain but not in Sweden, where the temperatures are similar to those of Britain during the MIS 7 period. Environment of boreal forest with summer temperatures between 14 °C and 18 °C and winter temperatures between 1 °C and −10 °C. The prey species includes elk, reindeer, roe deer, boar, hares, rabbit, and beaver. Competitors include the brown bear and lynx but the wolf is top carnivore. The wolves found in Sweden today are smaller in body size than MIS 5 wolves but larger than those of MIS 7 and MIS 3. The upper M1 and M2 length is longer than for MIS 7 and MIS 3 wolves, and the jaws deeper and broader, which indicates the ability to hunt and subdue large prey. However, the large molars retained a crushing ability and to process non-meat foods. These wolves live in boreal forests where small to medium game is hard to detect and labour-intensive to subdue, leading to an adaption for hunting larger game with higher reward. They are hypercarnivores similar to MIS 5 wolves but not with the same slicing ability.[151] |
During the Last Glacial Maximum 20,000 YBP, the Pleistocene steppe stretched across northern and central Eurasia and through Beringia into North America. The Pleistocene wolves of Beringia, and perhaps those across the steppe, were adapted to this habitat. Their tooth and skull morphology indicates that they specialized in preying on now-extinct Pleistocene megafauna, and their tooth wear indicates that their behavior was different to modern wolves.[106][141][160][161] This highlights the success of C. lupus as a species in adapting to different environmental conditions.[151] This gray wolf ecomorph became extinct at the end of the glaciation, along with the horse and other species on which it depended, and was replaced by wolves from southern North America. This indicates that specialized wolf ecomorphs can become extinct when their environment changes even though the habitat may still support other wolves.[141] Wolves went through a population bottleneck 20,000 YBP that coincides with the Last Glacial Maximum,[64][110][65][141] which indicates that many wolf populations may have gone extinct at the same time as the Beringian wolves.[141]
There are a small number of Canis remains that have been found at Goyet Cave, Belgium (36,500 YBP)[55] Razboinichya Cave, Russia (33,500 YBP)[162] Kostenki 8, Russia (33,500–26,500 YBP)[163] Predmosti, Czech Republic (31,000 YBP)[164] and Eliseevichi 1, Russia (17,000 YBP).[56] Based on cranial morphometric study of the characteristics thought to be associated with the domestication process, these have been proposed as early Paleolithic dogs.[163] These characteristics of shortened rostrum, tooth crowding, and absence or rotation of premolars have been documented in both ancient and modern wolves.[106][65][151][165][166][167] Rather than representing early dogs, these specimens may represent "a morphologically distinct local, now extinct, population of wolves".[65][168]
References
- ↑ Dawkins, William Boyd, Sir; Sanford, W. Ayshford; Reynolds, Sidney H. (1912). "British Pleistocene Hyænidæ, Ursidæ, Canidæ, and Mustelidæ". A Monograph of the British Pleistocene Mammalia. 2. London: Palaeontographical Society.
- 1 2 Sardella, Raffaele; Bertè, Davide; Iurino, Dawid Adam; Cherin, Marco; Tagliacozzo, Antonio (2014). "The wolf from Grotta Romanelli (Apulia, Italy) and its implications in the evolutionary history of Canis lupus in the Late Pleistocene of Southern Italy". Quaternary International. 328-329: 179–195. doi:10.1016/j.quaint.2013.11.016.
- ↑ Flynn, John J.; Wesley-Hunt, Gina D. (2005). "Phylogeny of the Carnivora: Basal Relationships Among the Carnivoramorphans, and Assessment of the Position of 'Miacoidea' Relative to Carnivora". Journal of Systematic Paleontology. 3: 1–28.
- ↑ "Eucyon davisi". Fossilworks. Retrieved 11 July 2016.
- 1 2 3 4 5 6 7 8 9 10 11 12 13 Wang, Xiaoming; Tedford, Richard H.; Dogs: Their Fossil Relatives and Evolutionary History. New York: Columbia University Press, 2008.
- 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 R.M. Nowak (2003). "Chapter 9 - Wolf evolution and taxonomy". In Mech, L. David; Boitani, Luigi. Wolves: Behaviour, Ecology and Conservation. University of Chicago Press. pp. 239–258. ISBN 978-0-226-51696-7.
- 1 2 3 4 5 6 7 8 9 10 11 12 Nowak, R. M. (1979). North American Quaternary Canis. 6. Monograph of the Museum of Natural History, University of Kansas. pp. 1–154. ISBN 978-0-89338-007-6.
- ↑ Wayne, R. K. (1995). "Red wolf: To conserve or not to conserve". In Macdonald, D.; Handoca, L. Canid News (PDF). 3. Newsletter of the IUCN/SSC Specialist Group. pp. 7–12.
- ↑ Hoffmeister, D. F., and W. W. Goodpaster. 1954. The mammals of the Huachuca Mountains, southeastern Arizona. Illinois Biological Monographs 24:1— 152.
- ↑ Lawrence, B.; Bossert, W. H. (1967). "Multiple character analysis of Canis lupus, latrans and niger". American Zoologist. 7: 223–232. doi:10.1093/icb/7.2.223. JSTOR 3881428.
- ↑ Lawrence, B.; Bossert, W. H. (1975). "Relationships of North American Canis shown by a multiple character analysis of selected populations". In Fox, M. W. Wild Canids: Their Systematics, Behavioral Ecology & Evolution. New York: Van Nostrand-Reinhold. ISBN 978-0-442-22430-1.
- 1 2 Martin, R. A.; Webb, S. D. (1974). "Late Pleistocene mammals from the Devil's Den fauna, Levy County". In Webb, S. D. Pleistocene Mammals of Florida. Gainesville: University Presses of Florida. pp. 114–145. ISBN 978-0-8130-0361-0.
- ↑ Webb, S. D. (1974). "Chronology of Florida Pleistocene mammals". Pleistocene Mammals of Florida. Gainesville: University Presses of Florida. pp. 5–31. ISBN 978-0-8130-0361-0.
- ↑ Wayne, R. K.; Jenks, S. M. (1991). "Mitochondrial DNA analysis supports extensive hybridization of the endangered red wolf (Canis rufus)". Nature. 351: 565–68. doi:10.1038/351565a0.
- ↑ "Canis lepophagus". Fossilworks. Retrieved 11 July 2016.
- 1 2 3 Kurten, B. (1974). "A History of Coyote-Like Dogs (Canidae, Mamalia)". Acta zoologica Fennica (140): 1–38.
- 1 2 3 4 5 Kurten, B.; Anderson, E. (1980). Pleistocene mammals of North America. New York: Columbia University Press. pp. 1–442. ISBN 978-0-231-03733-4.
- 1 2 Johnston, C. S. (1938). "Preliminary report on the vertebrate type locality of Cita Canyon and the description of an ancestral coyote". American Journal of Science. 5. 35 (209): 383–390. doi:10.2475/ajs.s5-35.209.383.
- 1 2 3 4 5 6 7 8 9 10 11 12 13 14 Tedford, Richard H.; Wang, Xiaoming; Taylor, Beryl E. (2009). "Phylogenetic Systematics of the North American Fossil Caninae (Carnivora: Canidae)" (PDF). Bulletin of the American Museum of Natural History. 325: 1–218. doi:10.1206/574.1.
- 1 2 Tedford, R.H. & Qiu, Z.-X., 1996 - A new canid genus from the Pliocene of Yushe, Shanxi Province - Vertebrata PalAsiatica 34 (1): 27-40
- ↑ Anderson, E. (1996). "A preliminary report on the Carnivora of Porcupine Cave, Park County, Colorado". In Churcher, C. S. R.; Stewart, K. M.; Seymour, K. L. Palaeoecology and palaeoenvironments of late Cenozoic mammals. Toronto: University of Toronto Press. pp. 259–282. ISBN 978-0-8020-0728-5.
- ↑ Albright, III, L. B. (2000). Biostratigraphy and Vertebrate Paleontology of the San Timoteo Badlands, Southern California. University of California Publications in Geological Sciences. 144. University of California Press. pp. 1–121. ISBN 978-0-520-91598-5.
- ↑ "Canis edwardii". Fossilworks. Retrieved 11 July 2016.
- ↑ Berta, A. (1995). "Fossil carnivores from the Leisey Shell Pits, Hillsborough County, Florida". In Hulbert, Jr., R. C.; Morgan, G. S.; Webb, S. D. Paleontology and geology of the Leisey Shell Pits, early Pleistocene of Florida. 37. Bulletin of the Florida Museum of Natural History. pp. 463–499.
- ↑ "Canis ambrusteri". Fossilworks. Retrieved 11 July 2016.
- ↑ Kurten, B. 1984: Geographic differentiation in the Rancholabrean dire wolf (Canis dirus Leidy) in North America. In Genoways, H. H. & Dawson, M. R. (eds.): Contributions in Quaternary Vertebrate Paleontology: A Volume in Memorial to John E. Guilday, 218–227. Carnegie Museum of Natural History Special Publication 8
- ↑ "Canis dirus". Fossilworks. Retrieved 11 July 2016.
- ↑ Goulet, G. D. (1993). Comparison of temporal and geographical skull variation among Nearctic, modern, Holocene, and late Pleistocene gray wolves (Canis lupus) and selected Canis (Master's Degree). Winnipeg: University of Manitoba. pp. 1–116.
- ↑ Berta, A. (1988). Quaternary evolution and biogeography of the large South American Canidae (Mammalia: Carnivora). 132. University of California Publications in Geological Sciences. pp. 1–49. ISBN 978-0-520-09960-9.
- ↑ Nowak, R. M.; Federoff, N. E. (2002). "The systematic status of the Italian wolf Canis lupus". Acta theriol. 47 (3): 333–338. doi:10.1007/BF03194151.
- 1 2 3 Kurtén, B. (1968). Pleistocene mammals of Europe. London: Weidenfeld and Nicolson. p. 317. ISBN 978-0-202-30953-8.
- ↑ Soergel, V. H. W. (1925). "Die Säugetierfauna des altdiluvialen Tonlagers von Jockgrim in der Plalz". Zeitschrift der Deutschen Geologischen Gesellschaft Abhandlungen (in German). 77: 405–438.
- 1 2 Sotnikova, M (2010). "Dispersal of the Canini (Mammalia, Canidae: Caninae) across Eurasia during the Late Miocene to Early Pleistocene". Quaternary International. 212 (2): 86–97. doi:10.1016/j.quaint.2009.06.008.
- ↑ Nowak, R. M. (1995). "Another look at wolf taxonomy". In Carbyn, L. H.; Fritts, S. H.; Seip, D. R. Ecology and Conservation of Wolves in a Changing World. Edmonton, Canada: Canadian Circumpolar Institute. pp. 375–397. ISBN 978-0-919058-92-7.
- ↑ Torre, D. (1967). "I cani Villafranchiani della Toscana". Palaeontographia Italica (in Italian). 63: 113–136.
- ↑ Torre, D. (1974). "Affinità dentali del cane della grotta di l'Escale". Rivista Italiana di Paleontologia e Stratigrafia (in Italian). 80: 147–156.
- ↑ Torre, D. (1979). "The Ruscinian and Villafranchian dogs of Europe". Bollettino della Società Paleontologica Italiana (in Italian). 18: 162–165.
- ↑ Martin, R. (1973). "Trois nouvelles espèces de Caninae (Canidae, Carnivora) des gisements Plio-Villafranchiens d'Europe. Documents des Laboratoires de Géologie de Lyon". Notes et Mémoires (in French). 57: 87–96.
- ↑ Sotnikova, M. (1989). "The carnivore mammals from the Pliocene to the early Pleistocene. Stratigraphic significance". Transactions of the Geological Institute of RAS. 440: 1–122.
- ↑ Rook, L. (1993). I cani dell’Eurasia dal Miocene superiore al Pleistocene medio (Ph.D.). Universities of Modena, Bologna, Firenze, Roma “La Sapienza”, Italy.
- ↑ Sotnikova, M. (2001). "Remains of Canidae from the Lower Pleistocene site of Untermassfeld". In Kahlke, R. D. Das Plestozan von Untermassfled bei Meiningen (Thuringen), part 2. 40. Romisch-Germanisches Zentralmuseum. pp. 607–632. ISBN 978-3-7749-3080-3.
- ↑ Brugal, J. P.; Boudadi-Maligne, M. (2011). "Quaternary small to large canids in Europe: taxonomic status and biochronological contribution". Quaternary International. 243: 171–182. doi:10.1016/j.quaint.2011.01.046.
- ↑ Cherin, M.; Bertè, D. F.; Sardella, R.; Rook, L. (2013). "Canis etruscus (Canidae, Mammalia) and its role in the faunal assemblage from Pantalla (Perugia, central Italy): comparison with the Late Villafranchian large carnivore guild of Italy". Bollettino della Società Paleontologica Italiana. 52 (1): 11–18. doi:10.4435/BSPI.2013.05.
- ↑ Cherin, Marco; Bertè, Davide; Rook, Lorenzo; Sardella, Raffaele (2014). "Re-Defining Canis etruscus (Canidae, Mammalia): A New Look into the Evolutionary History of Early Pleistocene Dogs Resulting from the Outstanding Fossil Record from Pantalla (Italy)". Journal of Mammalian Evolution. 21 (1): 95–110. doi:10.1007/s10914-013-9227-4.
- ↑ Lumley, H. de, Kahlke, H.D., Moigne, A.M., Moulle, P.E., 1988. Les faunes de grands mammifères de la grotte du Vallonet Roquebrune-Cap-Martin, Alpes-Maritimes. L’Anthropologie 92, 465–496.
- ↑ Soergel, W., 1928. Ein Kleiner Wolf aus dem Kiesen von Süssenborn. Zeitschrift der Deutschen Geologische Gesellschaft 80, 227–255.
- ↑ Garrido, Guiomar; Arribas, Alfonso (2008). "Canis accitanus nov. sp., a new small dog (Canidae, Carnivora, Mammalia) from the Fonelas P-1 Plio-Pleistocene site (Guadix basin, Granada, Spain)". Geobios. 41 (6): 751–761. doi:10.1016/j.geobios.2008.05.002.
- 1 2 Martínez-Navarro, Bienvenido; Belmaker, Miriam; Bar-Yosef, Ofer (2009). "The large carnivores from 'Ubeidiya (early Pleistocene, Israel): Biochronological and biogeographical implications". Journal of Human Evolution. 56 (5): 514–24. doi:10.1016/j.jhevol.2009.02.004. PMID 19427671.
- 1 2 New remains of Canis chihliensis (Mammalia, Carnivora) from Shanshenmiaozui, a lower Pleistocene site in Yangyuan, Hebei Tong, H.-w., N. Hu, and X. Wang. 2012. Vertebrata PalAsiatica 50(4):335-360
- 1 2 3 4 5 6 7 8 9 Koler-Matznick, Janice (2002). "The origin of the dog revisited" (PDF). Anthrozoos: A Multidisciplinary Journal of the Interactions of People & Animals. 15 (2): 98. doi:10.2752/089279302786992595.
- 1 2 Olsen, S. J. (1985). Origins of the domestic dog: the fossil record. Univ. of Arizona Press, Tucson, USA. pp. 88–89.
- ↑ Olsen S J, Olsen J W, Qi G Q, 1982. The position of Canis lupus variabilis, from Zhoukoudian, in the ancestral lineage of the domestic dog, Canis familiaris. Vert PalAsia, 20(3): 264-267(in Chinese with last page abstract in English)
- ↑ Lawrence, B. 1967. Early domestic dogs. Zeitschrift für Säugetierkunde 32: 44 – 59.
- ↑ Pei, W.C. (1934). The carnivora from locality 1 of Choukoutien. Palaeontologia Sinica, Series C, vol. 8, Fascicle 1. Geological Survey of China, Beijing. pp. 1–45.
- 1 2 3 Germonpré, Mietje; Sablin, Mikhail V.; Stevens, Rhiannon E.; Hedges, Robert E.M.; Hofreiter, Michael; Stiller, Mathias; Despre´s, Viviane R. (2009). "Fossil dogs and wolves from Palaeolithic sites in Belgium, the Ukraine and Russia: osteometry, ancient DNA and stable isotopes". Journal of Archaeological Science. 36 (2): 473–490. doi:10.1016/j.jas.2008.09.033.
- 1 2 Sablin, Mikhail V.; Khlopachev, Gennady A. (2002). "The Earliest Ice Age Dogs:Evidence from Eliseevichi I" (PDF). Wenner-Gren Foundation for Anthropological Research. Retrieved 10 January 2015.
- 1 2 3 4 5 6 7 8 9 10 11 Thalmann, O.; Shapiro, B.; Cui, P.; Schuenemann, V. J.; Sawyer, S. K.; Greenfield, D. L.; Germonpre, M. B.; Sablin, M. V.; Lopez-Giraldez, F.; Domingo-Roura, X.; Napierala, H.; Uerpmann, H.-P.; Loponte, D. M.; Acosta, A. A.; Giemsch, L.; Schmitz, R. W.; Worthington, B.; Buikstra, J. E.; Druzhkova, A.; Graphodatsky, A. S.; Ovodov, N. D.; Wahlberg, N.; Freedman, A. H.; Schweizer, R. M.; Koepfli, K.- P.; Leonard, J. A.; Meyer, M.; Krause, J.; Paabo, S.; et al. (2013). "Complete Mitochondrial Genomes of Ancient Canids Suggest a European Origin of Domestic Dogs". Science. 342 (6160): 871–4. doi:10.1126/science.1243650. PMID 24233726.
- 1 2 Morey, Darcy F.; Jeger, Rujana (2015). "Paleolithic dogs: Why sustained domestication then?". Journal of Archaeological Science: Reports. 3: 420. doi:10.1016/j.jasrep.2015.06.031.
- ↑ Wayne, Robert K. (1986). "Cranial Morphology of Domestic and Wild Canids: The Influence of Development on Morphological Change". Evolution. 40 (2): 243. doi:10.2307/2408805. JSTOR 2408805.
- ↑ Drake, Abby Grace; Klingenberg, Christian Peter (2010). "Large‐Scale Diversification of Skull Shape in Domestic Dogs: Disparity and Modularity". The American Naturalist. 175 (3): 289. doi:10.1086/650372. PMID 20095825.
- ↑ Roberts, Taryn; McGreevy, Paul; Valenzuela, Michael (2010). "Human Induced Rotation and Reorganization of the Brain of Domestic Dogs". PLoS ONE. 5 (7): e11946. doi:10.1371/journal.pone.0011946. PMID 20668685.
- ↑ Drake, Abby Grace, "Evolution and development of the skull morphology of canids: An investigation of morphological integration and heterochrony" (January 1, 2004). Doctoral Dissertations Available from Proquest. Paper AAI3136721. link
- ↑ Drake, Abby Grace (2011). "Dispelling dog dogma: An investigation of heterochrony in dogs using 3D geometric morphometric analysis of skull shape". Evolution & Development. 13 (2): 204. doi:10.1111/j.1525-142X.2011.00470.x.
- 1 2 3 4 5 6 7 8 9 10 11 12 Freedman, Adam H.; Gronau, Ilan; Schweizer, Rena M.; Ortega-Del Vecchyo, Diego; Han, Eunjung; Silva, Pedro M.; Galaverni, Marco; Fan, Zhenxin; Marx, Peter; Lorente-Galdos, Belen; Beale, Holly; Ramirez, Oscar; Hormozdiari, Farhad; Alkan, Can; Vilà, Carles; Squire, Kevin; Geffen, Eli; Kusak, Josip; Boyko, Adam R.; Parker, Heidi G.; Lee, Clarence; Tadigotla, Vasisht; Siepel, Adam; Bustamante, Carlos D.; Harkins, Timothy T.; Nelson, Stanley F.; Ostrander, Elaine A.; Marques-Bonet, Tomas; Wayne, Robert K.; et al. (2014). "Genome Sequencing Highlights the Dynamic Early History of Dogs". PLoS Genetics. 10 (1): e1004016. doi:10.1371/journal.pgen.1004016. PMC 3894170
 . PMID 24453982.
. PMID 24453982. - 1 2 3 4 5 6 7 Perri, Angela (2016). "A wolf in dog's clothing: Initial dog domestication and Pleistocene wolf variation". Journal of Archaeological Science. 68: 1–4. doi:10.1016/j.jas.2016.02.003.
- ↑ Rand, David M. (2001). "The Units of Selection on Mitochondrial DNA". Annual Review of Ecology and Systematics. 32: 415–448. doi:10.1146/annurev.ecolsys.32.081501.114109.
- 1 2 3 4 5 6 7 8 Robert K. Wayne, Jennifer A. Leonard, Carles Vila (2006). "Chapter 19:Genetic Analysis of Dog Domestication". In Melinda A. Zeder. Documenting Domestication:New Genetic and Archaeological Paradigms. University of California Press. pp. 279–295. ISBN 978-0-520-24638-6.
- ↑ Hofreiter, Michael; Serre, David; Poinar, Hendrik N.; Kuch, Melanie; Pääbo, Svante (2001). "Ancient DNA". Nature Reviews Genetics. 2 (5): 353–9. doi:10.1038/35072071. PMID 11331901.
- ↑ Li, Wen-Hsiung (1997). Molecular evolution. Sunderland, Massachusetts: Sinauer Associates. ISBN 978-0-87893-463-8.
- ↑ Pesole, G; Gissi, C; De Chirico, A; Saccone, C (1999). "Nucleotide substitution rate of mammalian mitochondrial genomes". Journal of Molecular Evolution. 48 (4): 427–34. doi:10.1007/pl00006487. PMID 10079281.
- ↑ Wan, Qiu-Hong; Wu, Hua; Fujihara, Tsutomu; Fang, Sheng-Guo (2003). "Which genetic marker for which conservation genetics issue?" (PDF). Electrophoresis. 25 (14): 2165. doi:10.1002/elps.200305922. PMID 15274000.
- ↑ Birky, C. William (2001). "The Inheritance of Genes in Mitochondria and Chloroplasts: Laws, Mechanisms, and Models". Annual Review of Genetics. 35: 125–48. doi:10.1146/annurev.genet.35.102401.090231. PMID 11700280.
- ↑ Avise, J. C. (1994). Molecular Markers, Natural History, and Evolution. New York: Chapman & Hall. ISBN 978-0-412-03781-8.
- ↑ Avise, J. C. (2000). Phylogeography: The History and Formation of Species. Cambridge, Massachusetts: Harvard University Press. ISBN 978-0-674-66638-2.
- ↑ Tedford, Richard H.; Wang, Xiaoming; Taylor, Beryl E. (2009). "Phylogenetic Systematics of the North American Fossil Caninae (Carnivora: Canidae)". Bulletin of the American Museum of Natural History. 325: 1–218. doi:10.1206/574.1.
- 1 2 3 4 5 6 7 Lindblad-Toh, K.; Wade, C. M.; Mikkelsen, T. S.; Karlsson, E. K.; Jaffe, D. B.; Kamal, M.; Clamp, M.; Chang, J. L.; Kulbokas, E. J.; Zody, M. C.; Mauceli, E.; Xie, X.; Breen, M.; Wayne, R. K.; Ostrander, E. A.; Ponting, C. P.; Galibert, F.; Smith, D. R.; Dejong, P. J.; Kirkness, E.; Alvarez, P.; Biagi, T.; Brockman, W.; Butler, J.; Chin, C. W.; Cook, A.; Cuff, J.; Daly, M. J.; Decaprio, D.; et al. (2005). "Genome sequence, comparative analysis and haplotype structure of the domestic dog". Nature. 438 (7069): 803–819. Bibcode:2005Natur.438..803L. doi:10.1038/nature04338. PMID 16341006.
- 1 2 3 4 5 6 Koepfli, K.-P.; Pollinger, J.; Godinho, R.; Robinson, J.; Lea, A.; Hendricks, S.; Schweizer, R. M.; Thalmann, O.; Silva, P.; Fan, Z.; Yurchenko, A. A.; Dobrynin, P.; Makunin, A.; Cahill, J. A.; Shapiro, B.; Álvares, F.; Brito, J. C.; Geffen, E.; Leonard, J. A.; Helgen, K. M.; Johnson, W. E.; O’Brien, S. J.; Van Valkenburgh, B.; Wayne, R. K. (2015-08-17). "Genome-wide Evidence Reveals that African and Eurasian Golden Jackals Are Distinct Species". Current Biology. 25 (16): 2158–65. doi:10.1016/j.cub.2015.06.060. PMID 26234211.
- 1 2 3 4 5 Vila, C. (1997). "Multiple and ancient origins of the domestic dog". Science. 276 (5319): 1687–9. doi:10.1126/science.276.5319.1687. PMID 9180076.
- 1 2 3 4 Wayne, R. & Ostrander, Elaine A. (1999). "Origin, genetic diversity, and genome structure of the domestic dog". BioEssays. 21 (3): 247–57. doi:10.1002/(SICI)1521-1878(199903)21:3<247::AID-BIES9>3.0.CO;2-Z. PMID 10333734.
- ↑ Boyko, A. (2009). "Complex population structure in African village dogs and its implications for inferring dog domestication history". Proceedings of the National Academy of Sciences. 106 (33): 13903–13908. doi:10.1073/pnas.0902129106. PMC 2728993
 . PMID 19666600.
. PMID 19666600. - ↑ Pang, J. (2009). "mtDNA data indicate a single origin for dogs south of Yangtze River, less than 16,300 years ago, from numerous wolves". Molecular Biology and Evolution. 26 (12): 2849–64. doi:10.1093/molbev/msp195. PMC 2775109
 . PMID 19723671.
. PMID 19723671. - 1 2 Cronin, M. A.; Canovas, A.; Bannasch, D. L.; Oberbauer, A. M.; Medrano, J. F. (2014). "Single Nucleotide Polymorphism (SNP) Variation of Wolves (Canis lupus) in Southeast Alaska and Comparison with Wolves, Dogs, and Coyotes in North America". Journal of Heredity. 106 (1): 26–36. doi:10.1093/jhered/esu075. PMID 25429025.
- ↑ Bardeleben, Carolyne; Moore, Rachael L.; Wayne, Robert K. (2005). "A molecular phylogeny of the Canidae based on six nuclear loci". Molecular Phylogenetics and Evolution. 37 (3): 815. doi:10.1016/j.ympev.2005.07.019. PMID 16213754.
- ↑ Gray, Melissa M; Sutter, Nathan B; Ostrander, Elaine A; Wayne, Robert K (2010). "The IGF1 small dog haplotype is derived from Middle Eastern grey wolves". BMC Biology. 8: 16. doi:10.1186/1741-7007-8-16. PMC 2837629
 . PMID 20181231.
. PMID 20181231. - ↑ Wayne, Robert K.; Vonholdt, Bridgett M. (2012). "Evolutionary genomics of dog domestication". Mammalian Genome. 23 (1–2): 3–18. doi:10.1007/s00335-011-9386-7. PMID 22270221.
- 1 2 3 4 5 6 7 8 9 10 11 12 13 14 Skoglund, P. (2015). "Ancient wolf genome reveals an early divergence of domestic dog ancestors and admixture into high-latitude breeds". Current Biology. 25 (11): 1515–9. doi:10.1016/j.cub.2015.04.019. PMID 26004765.
- 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 Fan, Zhenxin; Silva, Pedro; Gronau, Ilan; Wang, Shuoguo; Armero, Aitor Serres; Schweizer, Rena M.; Ramirez, Oscar; Pollinger, John; Galaverni, Marco; Ortega Del-Vecchyo, Diego; Du, Lianming; Zhang, Wenping; Zhang, Zhihe; Xing, Jinchuan; Vilà, Carles; Marques-Bonet, Tomas; Godinho, Raquel; Yue, Bisong; Wayne, Robert K. (2016). "Worldwide patterns of genomic variation and admixture in gray wolves". Genome Research. 26 (2): 163–73. doi:10.1101/gr.197517.115. PMID 26680994.
- ↑ Frantz, L. A. F.; Mullin, V. E.; Pionnier-Capitan, M.; Lebrasseur, O.; Ollivier, M.; Perri, A.; Linderholm, A.; Mattiangeli, V.; Teasdale, M. D.; Dimopoulos, E. A.; Tresset, A.; Duffraisse, M.; McCormick, F.; Bartosiewicz, L.; Gal, E.; Nyerges, E. A.; Sablin, M. V.; Brehard, S.; Mashkour, M.; b l Escu, A.; Gillet, B.; Hughes, S.; Chassaing, O.; Hitte, C.; Vigne, J.-D.; Dobney, K.; Hanni, C.; Bradley, D. G.; Larson, G. (2016). "Genomic and archaeological evidence suggest a dual origin of domestic dogs". Science. 352 (6290): 1228. doi:10.1126/science.aaf3161. PMID 27257259.
- 1 2 Wayne, R. (1993). "Molecular evolution of the dog family". Trends in Genetics. 9 (6): 218–24. doi:10.1016/0168-9525(93)90122-X. PMID 8337763.
- ↑ Wurster-Hill, D. H.; Centerwall, W. R. (1982). "The interrelationships of chromosome banding patterns in canids, mustelids, hyena, and felids". Cytogenetics and Cell Genetics. 34: 178–192. doi:10.1159/000131806. PMID 7151489.
- ↑ Gottelli, D.; Sillero-Zubiri, C.; Applebaum, G. D.; Roy, M. S.; Girman, D. J.; Garcia-Moreno, J.; Ostrander, E. A.; Wayne, R. K. (1994). "Molecular genetics of the most endangered canid: The Ethiopian wolf Canis simensis". Molecular Ecology. 3 (4): 301–12. doi:10.1111/j.1365-294X.1994.tb00070.x. PMID 7921357.
- ↑ Pocock, R. I. (1941). Fauna of British India: Mammals. 2. Taylor & Francis. pp. 146–163.
- ↑ Girman, D. J.; Vilà, C.; Geffen, E.; Creel, S.; Mills, M. G. L.; McNutt, J. W.; Ginsberg, J.; Kat, P. W.; Mamiya, K. H.; Wayne, R. K. (2001). "Patterns of population subdivision, gene flow and genetic variability in the African wild dog (Lycaon pictus)". Molecular Ecology. 10 (7): 1703–23. doi:10.1046/j.0962-1083.2001.01302.x. PMID 11472538.
- ↑ Wayne, R.K.; Meyer, A.; Lehman, N.; van Valkeburgh, B.; Kat, P.W.; Fuller, T.K.; Girman, D.; O'Brien, S.J. (1990). "Large sequence divergence among mitochondrial DNA genotypes within populations of eastern African black-backed jackals" (PDF). Proceedings of the National Academy of Sciences of the United States of America. 87: 1772–1776. doi:10.1073/pnas.87.5.1772. Retrieved 21 December 2011.
- ↑ Juliane Kaminski and Sarah Marshall-Pescini (2014). "Chapter 1 - The Social Dog:History and Evolution". The Social Dog:Behavior and Cognition. Elsevier. p. 4. ISBN 978-0-12407-931-1.
- 1 2 3 Aggarwal, R. K.; Kivisild, T.; Ramadevi, J.; Singh, L. (2007). "Mitochondrial DNA coding region sequences support the phylogenetic distinction of two Indian wolf species". Journal of Zoological Systematics and Evolutionary Research. 45 (2): 163–172. doi:10.1111/j.1439-0469.2006.00400.x.
- 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 Pilot, M.; et al. (2010). "Phylogeographic history of grey wolves in Europe". BMC Evolutionary Biology. 10: 104. doi:10.1186/1471-2148-10-104. PMC 2873414
 . PMID 20409299.
. PMID 20409299. - 1 2 3 4 5 6 Matsumura, Shuichi; Inoshima, Yasuo; Ishiguro, Naotaka (2014). "Reconstructing the colonization history of lost wolf lineages by the analysis of the mitochondrial genome". Molecular Phylogenetics and Evolution. 80: 105–12. doi:10.1016/j.ympev.2014.08.004. PMID 25132126.
- ↑ Cox, C. B.; Moore, Peter D.; Ladle, Richard (2016). Biogeography: An Ecological and Evolutionary Approach. Wiley-Blackwell. p. 106. ISBN 978-1-118-96858-1.
- ↑ Editorial Board (2012). Concise Dictionary of Science. New Delhi: V&S Publishers. p. 137. ISBN 978-93-81588-64-2.
- 1 2 Arora, Devender; Singh, Ajeet; Sharma, Vikrant; Bhaduria, Harvendra Singh; Patel, Ram Bahadur (2015). "Hgs Db: Haplogroups Database to understand migration and molecular risk assessment". Bioinformation. 11 (6): 272–5. doi:10.6026/97320630011272. PMC 4512000
 . PMID 26229286.
. PMID 26229286. - ↑ "Genetics Glossary". International Society of Genetic Genealogy. 2015. Retrieved 22 July 2016.
- ↑ Sharma, D. K.; Maldonado, J. E.; Jhala, Y. V.; Fleischer, R. C. (2004). "Ancient wolf lineages in India". Proceedings of the Royal Society B: Biological Sciences. 271 (Suppl 3): S1–S4. doi:10.1098/rsbl.2003.0071. PMC 1809981
 . PMID 15101402.
. PMID 15101402. - 1 2 3 Randi, Ettore (2011). "Genetics and conservation of wolves Canis lupus in Europe". Mammal Review. 41 (2): 99–111. doi:10.1111/j.1365-2907.2010.00176.x.
- ↑ Tamm, E.; Kivisild, T.; Reidla, M.; Metspalu, M.; Smith, D. G.; Mulligan, C. J.; Bravi, C. M.; Rickards, O.; Martinez-Labarga, C.; Khusnutdinova, E. K.; Fedorova, S. A.; Golubenko, M. V.; Stepanov, V. A.; Gubina, M. A.; Zhadanov, S. I.; Ossipova, L. P.; Damba, L.; Voevoda, M. I.; Dipierri, J. E.; Villems, R.; Malhi, R. S. (2007). Carter, Dee, ed. "Beringian Standstill and Spread of Native American Founders". PLoS ONE. 2 (9): e829. doi:10.1371/journal.pone.0000829. PMC 1952074
 . PMID 17786201.
. PMID 17786201. - 1 2 3 4 5 6 7 8 9 Leonard, J. A.; Vilà, C; Fox-Dobbs, K; Koch, P. L.; Wayne, R. K.; Van Valkenburgh, B (2007). "Megafaunal extinctions and the disappearance of a specialized wolf ecomorph" (PDF). Current Biology. 17 (13): 1146–50. doi:10.1016/j.cub.2007.05.072. PMID 17583509.
- 1 2 Hofreiter, Michael; Barnes, Ian (2010). "Diversity lost: Are all Holarctic large mammal species just relict populations?". BMC Biology. 8: 46. doi:10.1186/1741-7007-8-46. PMC 2858106
 . PMID 20409351.
. PMID 20409351. - ↑ Germonpré, Mietje; Sablin, Mikhail V.; Després, Viviane; Hofreiter, Michael; Lázničková-Galetová, Martina; Stevens, Rhiannon E.; Stiller, Mathias (2013). "Palaeolithic dogs and the early domestication of the wolf: A reply to the comments of Crockford and Kuzmin (2012)". Journal of Archaeological Science. 40: 786–792. doi:10.1016/j.jas.2012.06.016.
- 1 2 3 Hofreiter, Michael (2007). "Pleistocene Extinctions: Haunting the Survivors". Current Biology. 17 (15): R609–11. doi:10.1016/j.cub.2007.06.031. PMID 17686436.
- 1 2 3 Pilot, M; Greco, C; Vonholdt, B M; Jędrzejewska, B; Randi, E; Jędrzejewski, W; Sidorovich, V E; Ostrander, E A; Wayne, R K (2013). "Genome-wide signatures of population bottlenecks and diversifying selection in European wolves". Heredity. 112 (4): 428–42. doi:10.1038/hdy.2013.122. PMID 24346500.
- 1 2 3 Ersmark, Erik; Klütsch, Cornelya F. C.; Chan, Yvonne L.; Sinding, Mikkel-Holger S.; Fain, Steven R.; Illarionova, Natalia A.; Oskarsson, Mattias; Uhlén, Mathias; Zhang, Ya-Ping; Dalén, Love; Savolainen, Peter (2016). "From the Past to the Present: Wolf Phylogeography and Demographic History Based on the Mitochondrial Control Region". Frontiers in Ecology and Evolution. 4. doi:10.3389/fevo.2016.00134.
- 1 2 3 4 5 6 7 8 9 10 Koblmüller, Stephan; Vilà, Carles; Lorente-Galdos, Belen; Dabad, Marc; Ramirez, Oscar; Marques-Bonet, Tomas; Wayne, Robert K.; Leonard, Jennifer A. (2016). "Whole mitochondrial genomes illuminate ancient intercontinental dispersals of grey wolves (Canis lupus)". Journal of Biogeography. doi:10.1111/jbi.12765.
- ↑ Weckworth, Byron V.; Talbot, Sandra; Sage, George K.; Person, David K.; Cook, Joseph (2005). "A Signal for Independent Coastal and Continental histories among North American wolves". Molecular Ecology. 14 (4): 917. doi:10.1111/j.1365-294X.2005.02461.x. PMID 15773925.
- ↑ Weckworth, Byron V.; Talbot, Sandra L.; Cook, Joseph A. (2010). "Phylogeography of wolves (Canis lupus) in the Pacific Northwest". Journal of Mammalogy. 91 (2): 363. doi:10.1644/09-MAMM-A-036.1.
- ↑ Weckworth, Byron V.; Dawson, Natalie G.; Talbot, Sandra L.; Flamme, Melanie J.; Cook, Joseph A. (2011). "Going Coastal: Shared Evolutionary History between Coastal British Columbia and Southeast Alaska Wolves (Canis lupus)". PLoS ONE. 6 (5): e19582. doi:10.1371/journal.pone.0019582. PMID 21573241.
- 1 2 Vonholdt, B. M.; Cahill, J. A.; Fan, Z.; Gronau, I.; Robinson, J.; Pollinger, J. P.; Shapiro, B.; Wall, J.; Wayne, R. K. (2016). "Whole-genome sequence analysis shows that two endemic species of North American wolf are admixtures of the coyote and gray wolf". Science Advances. 2 (7): e1501714. doi:10.1126/sciadv.1501714.
- ↑ Abe, H. (1999). "Diversity and conservation of mammals of Japan". In Yokohata, Y.; Nakamura, S. Recent Advances in the Biology of Japanese Insectivore. Shōbara: Hiba Society of Natural History. pp. 89–104.
- ↑ Ishiguro, Naotaka; Inoshima, Yasuo; Shigehara, Nobuo; Ichikawa, Hideo; Kato, Masaru (2010). "Osteological and Genetic Analysis of the Extinct Ezo Wolf (Canis Lupus Hattai) from Hokkaido Island, Japan". Zoological Science. 27 (4): 320–4. doi:10.2108/zsj.27.320. PMID 20377350.
- ↑ Walker, Brett (2008). The Lost Wolves of Japan. University of Washington Press. p. 42.
- 1 2 3 Ishiguro, Naotaka; Inoshima, Yasuo; Shigehara, Nobuo (2009). "Mitochondrial DNA Analysis of the Japanese Wolf (Canis lupus hodophilax Temminck, 1839) and Comparison with Representative Wolf and Domestic Dog Haplotypes". Zoological Science. 26 (11): 765–70. doi:10.2108/zsj.26.765. PMID 19877836.
- 1 2 Pardi, Melissa I.; Smith, Felisa A. (2016). "Biotic responses of canids to the terminal Pleistocene megafauna extinction". Ecography. 39 (2): 141. doi:10.1111/ecog.01596.
- ↑ Witt, Kelsey E.; Judd, Kathleen; Kitchen, Andrew; Grier, Colin; Kohler, Timothy A.; Ortman, Scott G.; Kemp, Brian M.; Malhi, Ripan S. (2015). "DNA analysis of ancient dogs of the Americas: Identifying possible founding haplotypes and reconstructing population histories". Journal of Human Evolution. 79: 105–18. doi:10.1016/j.jhevol.2014.10.012. PMID 25532803.
- ↑ Wolf and Man. Evolution in Parallel. R.L. Hall and H.S. Sharp. Academic Press, New York, 1978
- ↑ Morell, Virginia (2016). "How do you save a wolf that's not really a wolf?". Science. 353 (6300). doi:10.1126/science.aag0699.
- ↑ Vila, C.; Amorim, I. R.; Leonard, J. A.; Posada, D.; Castroviejo, J.; Petrucci-Fonseca, F.; Crandall, K. A.; Ellegren, H.; Wayne, R. K. (1999). "Mitochondrial DNA phylogeography and population history of the grey wolf Canis lupus". Molecular Ecology. 8 (12): 2089–103. doi:10.1046/j.1365-294x.1999.00825.x. PMID 10632860.
- 1 2 Leonard, Jennifer A.; Vilà, Carles; Wayne, Robert K. (2004). "Fast Track: Legacy lost: Genetic variability and population size of extirpated US grey wolves (Canis lupus)". Molecular Ecology. 14 (1): 9–17. doi:10.1111/j.1365-294X.2004.02389.x. PMID 15643947.
- 1 2 Larson G, Bradley DG (2014). "How Much Is That in Dog Years? The Advent of Canine Population Genomics". PLOS Genetics. 10 (1): e1004093. doi:10.1371/journal.pgen.1004093.
- 1 2 Van Valkenburgh, Blaire; Hayward, Matthew W.; Ripple, William J.; Meloro, Carlo; Roth, V. Louise (2016). "The impact of large terrestrial carnivores on Pleistocene ecosystems". Proceedings of the National Academy of Sciences. 113 (4): 862–867. doi:10.1073/pnas.1502554112.
- ↑ Shipman, P. (2015). The Invaders:How humans and their dogs drove Neanderthals to extinction. Harvard University Press. ISBN 978-0-674-73676-4.
- ↑ Cagan, Alex; Blass, Torsten (2016). "Identification of genomic variants putatively targeted by selection during dog domestication". BMC Evolutionary Biology. 16. doi:10.1186/s12862-015-0579-7.
- ↑ Song, Jiao-Jiao; Wang, Wen-Zhi; Otecko, Newton O.; Peng, Min-Sheng; Zhang, Ya-Ping (2016). "Reconciling the conflicts between mitochondrial DNA haplogroup trees of Canis lupus". Forensic Science International: Genetics. 23: 83–85. doi:10.1016/j.fsigen.2016.03.008.
- 1 2 Gross, Michael (2015). "Are dogs just like us?". Current Biology. 25 (17): R733–6. doi:10.1016/j.cub.2015.08.018. PMID 26561653.
- ↑ "Ancient wolf genome pushes back dawn of the dog". Nature. 2015.
- ↑ Machugh, David E.; Larson, Greger; Orlando, Ludovic (2016). "Taming the Past: Ancient DNA and the Study of Animal Domestication". Annual Review of Animal Biosciences. 5. doi:10.1146/annurev-animal-022516-022747.
- ↑ Ersmark, E (2016). "Large carnivore population turnover and ecological change during the Late Quaternary". Stockholm University. Phd Thesis under J. Leonard and L. Dalen
- 1 2 Lee, E. (2015). "Ancient DNA analysis of the oldest canid species from the Siberian Arctic and genetic contribution to the domestic dog". PLoS ONE. 10 (5): e0125759. doi:10.1371/journal.pone.0125759. PMC 4446326
 . PMID 26018528.
. PMID 26018528. - ↑ Bocherens, Hervé; Drucker, Dorothée G.; Bonjean, Dominique; Bridault, Anne; Conard, Nicholas J.; Cupillard, Christophe; Germonpré, Mietje; Höneisen, Markus; Münzel, Susanne C.; Napierala, Hannes; Patou-Mathis, Marylène; Stephan, Elisabeth; Uerpmann, Hans-Peter; Ziegler, Reinhard (2011). "Isotopic evidence for dietary ecology of cave lion (Panthera spelaea) in North-Western Europe: Prey choice, competition and implications for extinction". Quaternary International. 245 (2): 249–261. doi:10.1016/j.quaint.2011.02.023.
- ↑ Yeakel, J. D.; Guimaraes, P. R.; Bocherens, H.; Koch, P. L. (2013). "The impact of climate change on the structure of Pleistocene food webs across the mammoth steppe". Proceedings of the Royal Society B: Biological Sciences. 280 (1762): 20130239. doi:10.1098/rspb.2013.0239.
- 1 2 Bocherens, Hervé (2015). "Isotopic tracking of large carnivore palaeoecology in the mammoth steppe". Quaternary Science Reviews. 117: 42–71. doi:10.1016/j.quascirev.2015.03.018.
- ↑ Dinnis, R.; Pate, A.; Reynolds, N. (2016). "Mid-to-late Marine Isotope Stage 3 mammal faunas of Britain: A new look". Proceedings of the Geologists' Association. 127 (4): 435. doi:10.1016/j.pgeola.2016.05.002.
- 1 2 3 4 5 6 7 8 9 10 11 12 13 14 Leonard, Jennifer (2014). "Ecology drives evolution in grey wolves" (PDF). Evolution Ecology Research. 16: 461–473.
- ↑ Vila, C.; Sundqvist, A.-K.; Flagstad, O.; Seddon, J.; Rnerfeldt, S. B.; Kojola, I.; Casulli, A.; Sand, H.; Wabakken, P.; Ellegren, H. (2003). "Rescue of a severely bottlenecked wolf (Canis lupus) population by a single immigrant". Proceedings of the Royal Society B: Biological Sciences. 270 (1510): 91–97. doi:10.1098/rspb.2002.2184.
- 1 2 3 Muñoz-Fuentes, Violeta; Darimont, Chris T.; Wayne, Robert K.; Paquet, Paul C.; Leonard, Jennifer A. (2009). "Ecological factors drive differentiation in wolves from British Columbia". Journal of Biogeography. 36 (8): 1516–1531. doi:10.1111/j.1365-2699.2008.02067.x.
- 1 2 3 Musiani, Marco; Leonard, Jennifer A.; Cluff, H. Dean; Gates, C. Cormack; Mariani, Stefano; Paquet, Paul C.; Vilà, Carles; Wayne, Robert K. (2007). "Differentiation of tundra/taiga and boreal coniferous forest wolves: Genetics, coat colour and association with migratory caribou". Molecular Ecology. 16 (19): 4149–70. doi:10.1111/j.1365-294X.2007.03458.x. PMID 17725575.
- ↑ Valière, Nathaniel; Fumagalli, Luca; Gielly, Ludovic; Miquel, Christian; Lequette, Benoît; Poulle, Marie-Lazarine; Weber, Jean-Marc; Arlettaz, Raphaël; Taberlet, Pierre (2003). "Long-distance wolf recolonization of France and Switzerland inferred from non-invasive genetic sampling over a period of 10 years". Animal Conservation. 6: 83–92. doi:10.1017/S1367943003003111.
- ↑ Fabbri, E.; Caniglia, R.; Kusak, J.; Galov, A.; Gomerčić, T.; Arbanasić, H.; Huber, D.; Randi, E. (2014). "Genetic structure of expanding wolf (Canis lupus) populations in Italy and Croatia, and the early steps of the recolonization of the Eastern Alps". Mammalian Biology - Zeitschrift für Säugetierkunde. 79 (2): 138–148. doi:10.1016/j.mambio.2013.10.002.
- ↑ Carmichael, L. E.; Nagy, J. A.; Larter, N. C.; Strobeck, C. (2001). "Prey specialization may influence patterns of gene flow in wolves of the Canadian Northwest". Molecular Ecology. 10 (12): 2787–98. doi:10.1046/j.0962-1083.2001.01408.x. PMID 11903892.
- ↑ Carmichael, L. E. (2006). Ecological Genetics of Northern Wolves and Arctic Foxes. Ph.D. Dissertation (PDF). University of Alberta.
- 1 2 Geffen, ELI; Anderson, Marti J.; Wayne, Robert K. (2004). "Climate and habitat barriers to dispersal in the highly mobile grey wolf". Molecular Ecology. 13 (8): 2481–90. doi:10.1111/j.1365-294X.2004.02244.x. PMID 15245420.
- ↑ Pilot, Malgorzata; Jedrzejewski, Wlodzimierz; Branicki, Wojciech; Sidorovich, Vadim E.; Jedrzejewska, Bogumila; Stachura, Krystyna; Funk, Stephan M. (2006). "Ecological factors influence population genetic structure of European grey wolves". Molecular Ecology. 15 (14): 4533–53. doi:10.1111/j.1365-294X.2006.03110.x. PMID 17107481.
- 1 2 3 4 5 6 7 8 Flower, Lucy O.H.; Schreve, Danielle C. (2014). "An investigation of palaeodietary variability in European Pleistocene canids". Quaternary Science Reviews. 96: 188–203. doi:10.1016/j.quascirev.2014.04.015.
- 1 2 Schweizer, Rena M.; Vonholdt, Bridgett M.; Harrigan, Ryan; Knowles, James C.; Musiani, Marco; Coltman, David; Novembre, John; Wayne, Robert K. (2016). "Genetic subdivision and candidate genes under selection in North American grey wolves". Molecular Ecology. 25 (1): 380–402. doi:10.1111/mec.13364. PMID 26333947.
- 1 2 Schweizer, Rena M.; Robinson, Jacqueline; Harrigan, Ryan; Silva, Pedro; Galverni, Marco; Musiani, Marco; Green, Richard E.; Novembre, John; Wayne, Robert K. (2016). "Targeted capture and resequencing of 1040 genes reveal environmentally driven functional variation in grey wolves". Molecular Ecology. 25 (1): 357–79. doi:10.1111/mec.13467. PMID 26562361.
- ↑ O'Keefe, F. Robin; Meachen, Julie; Fet, Elizabeth V.; Brannick, Alexandria (2013). "Ecological determinants of clinal morphological variation in the cranium of the North American gray wolf". Journal of Mammalogy. 94 (6): 1223–1236. doi:10.1644/13-MAMM-A-069.
- ↑ Carmichael, L. E.; Krizan, J.; Nagy, J. A.; Fuglei, E.; Dumond, M.; Johnson, D.; Veitch, A.; Berteaux, D.; Strobeck, C. (2007). "Historical and ecological determinants of genetic structure in arctic canids". Molecular Ecology. 16 (16): 3466–83. doi:10.1111/j.1365-294X.2007.03381.x. PMID 17688546.
- ↑ Lacombat, F.; Abbazzi, L.; Ferretti, M. P.; Martinez-Navarro, B.; Moulle, P.-E.; Palombo, M.-R.; Rook, L.; Turner, A.; Valli, A.M.-F. (2008). "New data on the early Villafranchian fauna from Vialette (Haute-Loire, France) based on the collection of the Crozatier Museum (Le Puy-en-Velay, Haute-Loire, France)". Quaternary International. 179 (1): 64–71. doi:10.1016/j.quaint.2007.09.005.
- ↑ Rook, L.; Torre, D. (1996). "The wolf-event in western Europe and the beginning of the Late Villafranchian". Neues Jahrbuch fur Geologie und Palaontologie Monatshefte. 8: 495–501.
- ↑ Gliozzi, E.; Abbazzi, L.; Argenti, A.; Azzaroli, A.; Caloi, L.; Capasso Barbato, L.; di Stefano, G.; Esu, D.; Ficcarelli, G.; Girotti, O.; Kotsakis, T.; Masini, F.; Mazza, P.; Mezzabotta, C.; Palombo, M. R.; Petronio, C.; Rook, L.; Sala, B.; Sardella, R.; Zanalda, E.; Torre, D. (1997). "Biochronology of selected mammals, Molluscs and ostracodes from the Middle Pliocene to the Late Pleistocene in Italy. The state of the art" (PDF). Rivista Italiana di Paleontologia e Stratigrafia. 103 (3): 369–388. doi:10.13130/2039-4942/5299.
- ↑ Currant, A. P. (1984). "The mammalian remains". In Green, H. S. Pontnewydd Cave: A Lower Palaeolithic Hominid Site in Wales: the First Report. Journal of Archaeological Science. 12. Amgueddfa Genedlaethol Cymru. pp. 177–181. doi:10.1016/0305-4403(85)90038-X. ISBN 978-0-7200-0282-9.
- ↑ Fox-Dobbs, Kena; Leonard, Jennifer A.; Koch, Paul L. (2008). "Pleistocene megafauna from eastern Beringia: Paleoecological and paleoenvironmental interpretations of stable carbon and nitrogen isotope and radiocarbon records". Palaeogeography, Palaeoclimatology, Palaeoecology. 261: 30–46. doi:10.1016/j.palaeo.2007.12.011.
- ↑ Baryshnikov, Gennady F.; Mol, Dick; Tikhonov, Alexei N (2009). "Finding of the Late Pleistocene carnivores in Taimyr Peninsula (Russia, Siberia) with paleoecological context" (PDF). Russian Journal of Theriology. Russian Journal of Theriology. 8 (2): 107–113. Retrieved December 23, 2014.
- ↑ Ovodov, N. (2011). "A 33,000-year-old incipient dog from the Altai Mountains of Siberia: Evidence of the earliest domestication disrupted by the Last Glacial Maximum". PLoS ONE. 6 (7): e22821. doi:10.1371/journal.pone.0022821. PMC 3145761
 . PMID 21829526.
. PMID 21829526. - 1 2 Germonpré, Mietje; Lázničková-Galetová, Martina; Losey, Robert J.; Räikkönen, Jannikke; Sablin, Mikhail V. (2015). "Large canids at the Gravettian Předmostí site, the Czech Republic: The mandible". Quaternary International. 359-360: 261–279. doi:10.1016/j.quaint.2014.07.012.
- ↑ Germonpré, Mietje; Laznickova-Galetova, Martina; Sablin, Mikhail V. (2012). "Palaeolithic dog skulls at the Gravettian Predmosti site, the Czech Republic". Journal of Archaeological Science. 39: 184–202. doi:10.1016/j.jas.2011.09.022.
- ↑ Boudadi-Maligne, M. (2010). "Les Canis pleistocenes du sud de la France: approche biosystematique, evolutive et biochronologique. Ph.D. dissertation" (PDF). Arch´eologie et Pr´ehistoire. Universitie Bordeaux (4126).
- ↑ Dimitrijević, V.; Vuković, S. (2015). "Was the Dog Locally Domesticated in the Danube Gorges? Morphometric Study of Dog Cranial Remains from Four Mesolithic-Early Neolithic Archaeological Sites by Comparison with Contemporary Wolves". International Journal of Osteoarchaeology. 25: 1–30. doi:10.1002/oa.2260.
- ↑ Germonpréa, Mietje; Sablinb, Mikhail V.; Stevensc, Rhiannon E.; Hedgesd, Robert E. M.; Hofreitere, Michael; Stillere, Mathias; Desprése, Viviane R. (2008). "Fossil dogs and wolves from Palaeolithic sites in Belgium, the Ukraine and Russia: osteometry, ancient DNA and stable isotopes". Journal of Archaeological Science. 36 (2): 473–490. doi:10.1016/j.jas.2008.09.033.
- ↑ Larson G (2012). "Rethinking dog domestication by integrating genetics, archeology, and biogeography". PNAS. 109 (23): 8878–8883. doi:10.1073/pnas.1203005109. PMC 3384140
 . PMID 22615366.
. PMID 22615366.