Integrin
Integrins are transmembrane receptors that are the bridges for cell-cell and cell-extracellular matrix (ECM) interactions. When triggered, integrins trigger chemical pathways to the interior (signal transduction), such as the chemical composition and mechanical status of the ECM. This results in a response (activation of transcription) like the regulation of the cell cycle, cell shape, and/or motility; or new receptors being added to the cell membrane. This allows rapid and flexible responses to events at the cell surface, for example to signal platelets to initiate an interaction with coagulation factors.
There are several types of integrins, and a cell may have several types on its surface. Integrins are found in all metazoa.[3]
Integrins work alongside other receptors such as cadherins, the immunoglobulin superfamily cell adhesion molecules, selectins and syndecans to mediate cell–cell and cell–matrix interaction. Ligands for integrins include fibronectin, vitronectin, collagen, and laminin.
Structure
Integrins are obligate heterodimers, meaning that they have two different chains: the α (alpha) and β (beta) subunits. In mammals, there are eighteen α and eight β subunits, in Drosophila five α and two β subunits, and in Caenorhabditis nematodes two α subunits and one β subunit.[4] The α and β subunits each penetrate the plasma membrane and possess small cytoplasmic domains.[5]
| gene | protein | synonyms |
|---|---|---|
| ITGA1 | CD49a | VLA1 |
| ITGA2 | CD49b | VLA2 |
| ITGA3 | CD49c | VLA3 |
| ITGA4 | CD49d | VLA4 |
| ITGA5 | CD49e | VLA5 |
| ITGA6 | CD49f | VLA6 |
| ITGA7 | ITGA7 | FLJ25220 |
| ITGA8 | ITGA8 | |
| ITGA9 | ITGA9 | RLC |
| ITGA10 | ITGA10 | |
| ITGA11 | ITGA11 | HsT18964 |
| ITGAD | CD11D | FLJ39841 |
| ITGAE | CD103 | HUMINAE |
| ITGAL | CD11a | LFA1A |
| ITGAM | CD11b | MAC-1 |
| ITGAV | CD51 | VNRA, MSK8 |
| ITGA2B | CD41 | GPIIb |
| ITGAX | CD11c |
| gene | protein | synonyms |
|---|---|---|
| ITGB1 | CD29 | FNRB, MSK12, MDF2 |
| ITGB2 | CD18 | LFA-1, MAC-1, MFI7 |
| ITGB3 | CD61 | GP3A, GPIIIa |
| ITGB4 | CD104 | |
| ITGB5 | ITGB5 | FLJ26658 |
| ITGB6 | ITGB6 | |
| ITGB7 | ITGB7 | |
| ITGB8 | ITGB8 |
Variants of some of the subunits are formed by differential RNA splicing; for example, four variants of the beta-1 subunit exist. Through different combinations of the α and β subunits, around 24 unique integrins are generated.[6] (See Vertebrate integrins below)
Integrin subunits span the cell membrane and have short cytoplasmic domains of 40–70 amino acids. The exception is the beta-4 subunit, which has a cytoplasmic domain of 1088 amino acids, one of the largest known cytoplasmic domains of any membrane protein. Outside the cell membrane, the α and β chains lie close together along a length of about 23 nm; the final 5 nm N-termini of each chain forms a ligand-binding region for the ECM. They have been compared to lobster claws, although the don't actually "pinch" their ligand, they chemically interact with it at the insides of the "tips" of their "pinchers".
The molecular mass of the integrin subunits can vary from 90 kDa to 160 kDa. Beta subunits have four cysteine-rich repeated sequences. Both α and β subunits bind several divalent cations. The role of divalent cations in the α subunit is unknown, but may stabilize the folds of the protein. The cations in the β subunits are more interesting: they are directly involved in coordinating at least some of the ligands that integrins bind.
There are various ways of categorizing the integrins. For example, a subset of the α chains has an additional structural element (or "domain") inserted toward the N-terminal, the alpha-A domain (so called because it has a similar structure to the A-domains found in the protein von Willebrand factor; it is also termed the α-I domain). Integrins carrying this domain either bind to collagens (e.g. integrins α1 β1, and α2 β1), or act as cell-cell adhesion molecules (integrins of the β2 family). This α-I domain is the binding site for ligands of such integrins. Those integrins that don't carry this inserted domain also have an A-domain in their ligand binding site, but this A-domain is found on the β subunit.
In both cases, the A-domains carry up to three divalent cation binding sites. One is permanently occupied in physiological concentrations of divalent cations, and carries either a calcium or magnesium ion, the principal divalent cations in blood at median concentrations of 1.4 mM (calcium) and 0.8 mM (magnesium). The other two sites become occupied by cations when ligands bind—at least for those ligands involving an acidic amino acid in their interaction sites. An acidic amino acid features in the integrin-interaction site of many ECM proteins, for example as part of the amino acid sequence Arginine-Glycine-Aspartic acid ("RGD" in the one-letter amino acid code).
Structure
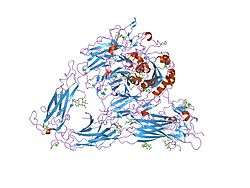
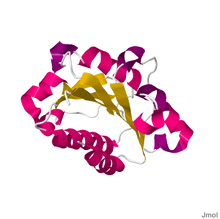
Despite many years of effort, discovering the high-resolution structure of integrins proved to be challenging, as membrane proteins are classically difficult to purify, and as integrins are large, complex and linked to many sugar trees ("highly glycosylated"). Low-resolution images of detergent extracts of intact integrin GPIIbIIIa, obtained using electron microscopy, and even data from indirect techniques that investigate the solution properties of integrins using ultracentrifugation and light scattering, were combined with fragmentary high-resolution crystallographic or NMR data from single or paired domains of single integrin chains, and molecular models postulated for the rest of the chains.
Despite these wide-ranging efforts, the X-ray crystal structure obtained for the complete extracellular region of one integrin, αvβ3, determined in 2001 by the laboratory of Dr. M. Amin Arnaout, MD, at Massachusetts General Hospital and Harvard Medical School, was a surprise.[7] It showed the molecule to be folded into an inverted V-shape that potentially brings the ligand-binding sites close to the cell membrane. Perhaps more importantly, the crystal structure was also obtained for the same integrin bound to a small ligand containing the RGD-sequence, the drug cilengitide.[8] As detailed above, this finally revealed why divalent cations (in the A-domains) are critical for RGD-ligand binding to integrins. The interaction of such sequences with integrins is believed to be a primary switch by which ECM exerts its effects on cell behaviour.
The structure poses many questions, especially regarding ligand binding and signal transduction. The ligand binding site is directed towards the C-terminal of the integrin, the region where the molecule emerges from the cell membrane. If it emerges orthogonally from the membrane, the ligand binding site would apparently be obstructed, especially as integrin ligands are typically massive and well cross-linked components of the ECM. In fact, little is known about the angle that membrane proteins subtend to the plane of the membrane; this is a problem difficult to address with available technologies. The default assumption is that they emerge rather like little lollipops, but the evidence for this sweet supposition is noticeable by its absence. The integrin structure has drawn attention to this problem, which may have general implications for how membrane proteins work. It appears that the integrin transmembrane helices are tilted (see "Activation" below), which hints that the extracellular chains may also not be orthogonal with respect to the membrane surface.
Although the crystal structure changed surprisingly little after binding to cilengitide, the current hypothesis is that integrin function involves changes in shape to move the ligand-binding site into a more accessible position, away from the cell surface, and this shape change also triggers intracellular signaling. There is a wide body of cell-biological and biochemical literature that supports this view. Perhaps the most convincing evidence involves the use of antibodies that only recognize integrins when they have bound to their ligands, or are activated. As the "footprint" that an antibody makes on its binding target is roughly a circle about 3 nm in diameter, the resolution of this technique is low. Nevertheless, these so-called LIBS (Ligand-Induced-Binding-Sites) antibodies unequivocally show that dramatic changes in integrin shape routinely occur. However, how the changes detected with antibodies look on the structure is still unknown.
Activation
When released into the cell membrane, newly synthesized integrin dimers are speculated to be found in the same "bent" conformation revealed by the structural studies described above. One school of thought claims that this bent form prevents them from interacting with their ligands, although bent forms can predominate in high-resolution EM structures of integrin bound to an ECM ligand. Therefore, at least in biochemical experiments, integrin dimers must apparently not be 'unbent' in order to prime them and allow their binding to the ECM. In cells, the priming is accomplished by a protein talin, which binds to the β tail of the integrin dimer and changes its conformation.[9][10] The α and β integrin chains are both class-I transmembrane proteins: they pass the plasma membrane as single transmembrane alpha-helices. Unfortunately, the helices are too long, and recent studies suggest that, for integrin gpIIbIIIa, they are tilted with respect both to one another and to the plane of the membrane. Talin binding alters the angle of tilt of the β3 chain transmembrane helix in model systems and this may reflect a stage in the process of inside-out signalling which primes integrins.[11] Moreover, talin proteins are able to dimerize[12] and thus are thought to intervene in the clustering of integrin dimers which leads to the formation of a focal adhesion. Recently, the Kindlin-1 and Kindlin-2 proteins have also been found to interact with integrin and activate it.[13]
Function
Integrins have two main functions:-
- Attachment of the cell to the ECM
- Signal transduction from the ECM to the cell
However, they are also involved in a wide range of other biological activities, including immune patrolling, cell migration, and binding to cells by certain viruses, such as adenovirus, echovirus, hantavirus, and foot and mouth disease viruses.
A prominent function of the integrins is seen in the molecule GPIIbIIIa, an integrin on the surface of blood platelets (thrombocytes) responsible for attachment to fibrin within a developing blood clot. This molecule dramatically increases its binding affinity for fibrin/fibrinogen through association of platelets with exposed collagens in the wound site. Upon association of platelets with collagen, GPIIbIIIa changes shape, allowing it to bind to fibrin and other blood components to form the clot matrix and stop blood loss.
Attachment of cell to the ECM
Integrins couple the ECM outside a cell to the cytoskeleton (in particular, the microfilaments) inside the cell. Which ligand in the ECM the integrin can bind to is defined by which α and β subunits the integrin is made of. Among the ligands of integrins are fibronectin, vitronectin, collagen, and laminin. The connection between the cell and the ECM may help the cell to endure pulling forces without being ripped out of the ECM. The ability of a cell to create this kind of bond is also of vital importance in ontogeny.
Cell attachment to the ECM is a basic requirement to build a multicellular organism. Integrins are not simply hooks, but give the cell critical signals about the nature of its surroundings. Together with signals arising from receptors for soluble growth factors like VEGF, EGF, and many others, they enforce a cellular decision on what biological action to take, be it attachment, movement, death, or differentiation. Thus integrins lie at the heart of many cellular biological processes. The attachment of the cell takes place through formation of cell adhesion complexes, which consist of integrins and many cytoplasmic proteins, such as talin, vinculin, paxillin, and alpha-actinin. These act by regulating kinases such as FAK (focal adhesion kinase) and Src kinase family members to phosphorylate substrates such as p130CAS thereby recruiting signaling adaptors such as CRK. These adhesion complexes attach to the actin cytoskeleton. The integrins thus serve to link two networks across the plasma membrane: the extracellular ECM and the intracellular actin filamentous system. Integrin alpha6beta4 is an exception: it links to the keratin intermediate filament system in epithelial cells.
Focal adhesions are large molecular complexes, which are generated following interaction of integrins with ECM, then their clustering. The clusters likely provide sufficient intracellular binding sites to permit the formation of stable signaling complexes on the cytoplasmic side of the cell membrane. So the focal adhesions contain integrin ligand, integrin molecule, and associate plaque proteins. Binding is propelled by changes in free energy.[14] As previously stated, these complexes connect the extracellular matrix to actin bundles. Cryo-electron tomography reveals that the adhesion contains particles on the cell membrane with diameter of 25 +/- 5 nm and spaced at approximately 45 nm.[15] Treatment with Rho-kinase inhibitor Y-27632 reduces the size of the particle, and it is extremely mechanosensitive.[16]
One important function of integrins on cells in tissue culture is their role in cell migration. Cells adhere to a substrate through their integrins. During movement, the cell makes new attachments to the substrate at its front and concurrently releases those at its rear. When released from the substrate, integrin molecules are taken back into the cell by endocytosis; they are transported through the cell to its front by the endocytic cycle, where they are added back to the surface. In this way they are cycled for reuse, enabling the cell to make fresh attachments at its leading front. It is not yet clear whether cell migration in tissue culture is an artefact of integrin processing, or whether such integrin-dependent cell migration also occurs in living organisms.
Signal transduction
Integrins play an important role in cell signaling by modulating the cell signaling pathways of transmembrane protein kinases such as receptor tyrosine kinases (RTK). While the interaction between integrin and receptor tyrosine kinases originally was thought of as uni-directional and supportive, recent studies indicate that integrins have additional, multi-faceted roles in cell signaling. Integrins can regulate the receptor tyrosine kinase signaling by recruiting specific adaptors to the plasma membrane. For example, β1c integrin recruits Gab1/Shp2 and presents Shp2 to IGF1R, resulting in dephosphorylation of the receptor.[17] In a reverse direction, when a receptor tyrosine kinase is activated, integrins co-localise at focal adhesion with the receptor tyrosine kinases and their associated signaling molecules.
The repertoire of integrins expressed on a particular cell can specify the signaling pathway due to the differential binding affinity of ECM ligands for the integrins. The tissue stiffness and matrix composition can initiate specific signaling pathways regulating cell behavior. Clustering and activation of the integrins/actin complexes strengthen the focal adhesion interaction and initiate the framework for cell signaling through assembly of adhesomes.[18]
Depending on the integrin's regulatory impact on specific receptor tyrosine kinases, the cell can experience:
- cell growth,
- cell division,
- cell survival,
- cellular differentiation, and
- apoptosis (programmed cell death).
Knowledge of the relationship between integrins and receptor tyrosine kinase has laid a foundation for new approaches to cancer therapy. Specifically, targeting integrins associated with RTKs is an emerging approach for inhibiting angiogenesis.[19]
Vertebrate integrins
The following are 16 of the ~24 integrins found in vertebrates:
| Name | Synonyms | Distribution | Ligands |
| α1β1 | VLA-1 | Many | Collagens, laminins[20] |
| α2β1 | VLA-2 | Many | Collagens, laminins[20] |
| α3β1 | VLA-3 | Many | Laminin-5 |
| α4β1 | VLA-4[20] | Hematopoietic cells | Fibronectin, VCAM-1[20] |
| α5β1 | VLA-5; fibronectin receptor | widespread | fibronectin[20] and proteinases |
| α6β1 | VLA-6; laminin receptor | widespread | laminins |
| α7β1 | muscle, glioma | laminins | |
| αLβ2 | LFA-1[20] | T-lymphocytes | ICAM-1, ICAM-2[20] |
| αMβ2 | Mac-1, CR3[20] | Neutrophils and monocytes | Serum proteins, ICAM-1[20] |
| αIIbβ3 | Fibrinogen receptor; gpIIbIIIa[21] | Platelets[20] | fibrinogen, fibronectin[20] |
| αVβ1 | ocular melanoma; neurological tumors | vitronectin; fibrinogen | |
| αVβ3 | vitronectin receptor[22] | activated endothelial cells, melanoma, glioblastoma | vitronectin,[22] fibronectin, fibrinogen, osteopontin, Cyr61, thyroxine,[23] TETRAC |
| αVβ5 | widespread, esp. fibroblasts, epithelial cells | vitronectin and adenovirus | |
| αVβ6 | proliferating epithelia, esp. lung and mammary gland | fibronectin; TGFβ1+3 | |
| αVβ8 | neural tissue; peripheral nerve | fibronectin; TGFβ1+3 | |
| α6β4 | Epithelial cells[20] | Laminin[20] |
Beta-1 integrins interact with many alpha integrin chains. Gene knockouts of integrins in mice are not always lethal, which suggests that during embryonal development, one integrin may substitute its function for another in order to allow survival. Some integrins are on the cell surface in an inactive state, and can be rapidly primed, or put into a state capable of binding their ligands, by cytokines. Integrins can assume several different well-defined shapes or "conformational states". Once primed, the conformational state changes to stimulate ligand binding, which then activates the receptors — also by inducing a shape change — to trigger outside-in signal transduction.
References
- ↑ Xiong JP, Stehle T, Diefenbach B, Zhang R, Dunker R, Scott DL, Joachimiak A, Goodman SL, Arnaout MA (2001). "Crystal structure of the extracellular segment of integrin alpha Vbeta3". Science. 294 (5541): 339–45. doi:10.1126/science.1064535. PMC 2885948
 . PMID 11546839.
. PMID 11546839. - ↑ Sauer FG, Fütterer K, Pinkner JS, Dodson KW, Hultgren SJ, Waksman G. "Structural basis of chaperone function and pilus biogenesis". Science. 285 (5430): 1058–61. doi:10.1126/science.285.5430.1058. PMID 10446050.
- ↑ Hynes RO (2002). "Integrins: bidirectional, allosteric signaling machines". Cell. 110 (6): 673–87. doi:10.1016/s0092-8674(02)00971-6. PMID 12297042.
- ↑ Humphries MJ (2000). "Integrin structure". Biochem. Soc. Trans. 28 (4): 311–339. doi:10.1042/0300-5127:0280311. PMID 10961914.
- ↑ Nermut MV, Green NM, Eason P, Yamada SS, Yamada KM (December 1988). "Electron microscopy and structural model of human fibronectin receptor". EMBO J. 7 (13): 4093–9. PMC 455118
 . PMID 2977331.
. PMID 2977331. - ↑ Hynes RO (2002). "Integrins: bidirectional, allosteric signaling machines". Cell. 110 (6): 673–87. doi:10.1016/S0092-8674(02)00971-6. PMID 12297042.
- ↑ Xiong JP, Stehle T, Diefenbach B, Zhang R, Dunker R, Scott DL, Joachimiak A, Goodman SL, Arnaout MA (2001). "Crystal structure of the extracellular segment of integrin αvβ3". Science. 294 (5541): 339–345. doi:10.1126/science.1064535. PMC 2885948
 . PMID 11546839.
. PMID 11546839. - ↑ Smith JW (2003). "Cilengitide Merck". Curr Opin Investig Drugs. 4 (6): 741–5. PMID 12901235.
- ↑ Calderwood DA (June 2004). "Talin controls integrin activation". Biochem. Soc. Trans. 32 (Pt3): 434–7. doi:10.1042/BST0320434. PMID 15157154.
- ↑ Calderwood DA, Zent R, Grant R, Rees DJ, Hynes RO, Ginsberg MH (October 1999). "The Talin head domain binds to integrin beta subunit cytoplasmic tails and regulates integrin activation". J. Biol. Chem. 274 (40): 28071–4. doi:10.1074/jbc.274.40.28071. PMID 10497155.
- ↑ Shattil SJ, Kim C, Ginsberg MH (2010). "The final steps of integrin activation: the end game". Nature Reviews Molecular Cell Biology. 11 (4): 288–300. doi:10.1038/nrm2871. PMID 20308986.
- ↑ Goldmann WH, Bremer A, Häner M, Aebi U, Isenberg G (1994). "Native talin is a dumbbell-shaped homodimer when it interacts with actin". J. Struct. Biol. 112 (1): 3–10. doi:10.1006/jsbi.1994.1002. PMID 8031639.
- ↑ Harburger DS, Bouaouina M, Calderwood DA (April 2009). "Kindlin-1 and −2 directly bind the C-terminal region of beta integrin cytoplasmic tails and exert integrin-specific activation effects". J. Biol. Chem. 284 (17): 11485–97. doi:10.1074/jbc.M809233200. PMC 2670154
 . PMID 19240021.
. PMID 19240021. - ↑ Olberding JE, Thouless MD, Arruda EM, Garikipati K (2010). Buehler MJ, ed. "The non-equilibrium thermodynamics and kinetics of focal adhesion dynamics". PLoS ONE. 5 (8): e12043. doi:10.1371/journal.pone.0012043. PMC 2923603
 . PMID 20805876.
. PMID 20805876. - ↑ Patla I, Volberg T, Elad N, Hirschfeld-Warneken V, Grashoff C, Fässler R, Spatz JP, Geiger B, Medalia O (September 2010). "Dissecting the molecular architecture of integrin adhesion sites by cryo-electron tomography". Nat. Cell Biol. 12 (9): 909–15. doi:10.1038/ncb2095. PMID 20694000.
- ↑ Gullingsrud J, Sotomayor M. "Mechanosensitive channels". Theoretical and Computational Biophysics Group, Beckman Institute for Advanced Science and Technology: University of Illinois at Urbana-Champaign.
- ↑ Goel HL, Breen M, Zhang J, Das I, Aznavoorian-Cheshire S, Greenberg NM, Elgavish A, Languino LR (1 August 2005). "1A Integrin Expression Is Required for Type 1 Insulin-Like Growth Factor Receptor Mitogenic and Transforming Activities and Localization to Focal Contacts". Cancer Research. 65 (15): 6692–6700. doi:10.1158/0008-5472.CAN-04-4315. PMID 16061650.
- ↑ Kim SH, Turnbull J, Guimond S (2011). "Extracellular matrix and cell signalling: the dynamic cooperation of integrin, proteoglycan and growth factor receptor". J. Endocrinol. 209 (2): 139–51. doi:10.1530/JOE-10-0377. PMID 21307119.
- ↑ Carbonell WS, DeLay M, Jahangiri A, Park CC, Aghi MK (3 May 2013). "1 Integrin Targeting Potentiates Antiangiogenic Therapy and Inhibits the Growth of Bevacizumab-Resistant Glioblastoma". Cancer Research. 73 (10): 3145–3154. doi:10.1158/0008-5472.CAN-13-0011. PMC 4040366
 . PMID 23644530.
. PMID 23644530. - 1 2 3 4 5 6 7 8 9 10 11 12 13 Krieger M, Scott MP, Matsudaira PT, Lodish HF, Darnell JE, Zipursky L, Kaiser C, Berk A (2004). Molecular cell biology (fifth ed.). New York: W.H. Freeman and CO. ISBN 0-7167-4366-3.
- ↑ Elangbam CS, Qualls CW, Dahlgren RR (1997). "Cell adhesion molecules--update" (PDF). Vet. Pathol. 34 (1): 61–73. doi:10.1177/030098589703400113. PMID 9150551.
- 1 2 Hermann P, Armant M, Brown E, Rubio M, Ishihara H, Ulrich D, Caspary RG, Lindberg FP, Armitage R, Maliszewski C, Delespesse G, Sarfati M (February 1999). "The vitronectin receptor and its associated CD47 molecule mediates proinflammatory cytokine synthesis in human monocytes by interaction with soluble CD23". J. Cell Biol. 144 (4): 767–75. doi:10.1083/jcb.144.4.767. PMC 2132927
 . PMID 10037797.
. PMID 10037797. - ↑ Bergh, JJ; Lin, HY; Lansing, L; Mohamed, SN; Davis, FB; Mousa, S; Davis, PJ (July 2005). "Integrin alphaVbeta3 contains a cell surface receptor site for thyroid hormone that is linked to activation of mitogen-activated protein kinase and induction of angiogenesis.". Endocrinology. 146 (7): 2864–71. doi:10.1210/en.2005-0102. PMID 15802494.
External links
![]() Media related to Integrins at Wikimedia Commons
Media related to Integrins at Wikimedia Commons
- MBInfo - Integrin-Mediated Signalling
- MBInfo - Integrin Activation
- Eukaryotic Linear Motif resource motif class LIG_RGD
- Eukaryotic Linear Motif resource motif class LIG_Integrin_isoDGR_1
- The Integrin Protein
- Talin substrate for calpain – PMAP The Proteolysis Map animation.
- Integrins at the US National Library of Medicine Medical Subject Headings (MeSH)