Purine metabolism
Purine metabolism refers to the metabolic pathways to synthesize and break down purines that are present in many organisms.
Biosynthesis
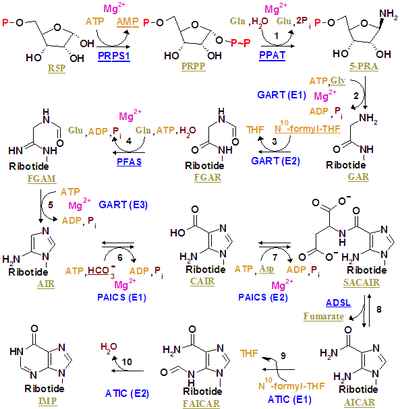
Purines are biologically synthesized as nucleotides and in particular as ribotides, i.e. bases attached to ribose 5-phosphate. A key regulatory step is the production of 5-phospho-α-D-ribosyl 1-pyrophosphate (PRPP) by ribose phosphate pyrophosphokinase, which is activated by inorganic phosphate and inactivated by purine ribonucleotides. It is not the committed step to purine synthesis because PRPP is also used in pyrimidine synthesis and salvage pathways.
The first committed step is the reaction of PRPP, glutamine and water to 5'-phosphoribosylamine (PRA), glutamate, and pyrophosphate - catalyzed by amidophosphoribosyltransferase, which is activated by PRPP and inhibited by AMP, GMP and IMP.
PRPP + L-Glutamine + H2O → PRA + L-Glutamate + PPi
In the second step react PRA, glycine and ATP to create GAR, ADP, and pyrophosphate - catalyzed by phosphoribosylamine—glycine ligase (GAR synthetase). Due to the chemical lability of PRA, which has a half-life of 38 seconds at PH 7.5 and 37 °C, researchers have suggested that the compound is channeled from amidophosphoribosyltransferase to GAR synthetase in vivo.[1]
PRA + Glycine + ATP → GAR + ADP + Pi
The third is catalyzed by phosphoribosylglycinamide formyltransferase.
The fourth is catalyzed by phosphoribosylformylglycinamidine synthase.
fGAR + L-Glutamine + ATP → fGAM + L-Glutamate + ADP + Pi
The fifth is catalyzed by AIR synthetase (FGAM cyclase).
fGAM + ATP → AIR + ADP + Pi + H2O
The sixth is catalyzed by phosphoribosylaminoimidazole carboxylase.
The seventh is catalyzed by phosphoribosylaminoimidazolesuccinocarboxamide synthase.
CAIR + L-Aspartate + ATP → SAICAR + ADP + Pi
The eight is catalyzed by adenylosuccinate lyase.
The ninth is catalyzed by phosphoribosylaminoimidazolecarboxamide formyltransferase.
AICAR + fTHF → FAICAR + THF
The last step is catalyzed by Inosine monophosphate synthase.
FAICAR → IMP + H2O
In eukaryotes the second, third, and fifth step are catalyzed by trifunctional purine biosynthetic protein adenosine-3, which is encoded by the GART gene.
Both ninth and tenth step are accomplished by a single protein named Bifunctional purine biosynthesis protein PURH, encoded by the ATIC gene.
Both adenine and guanine are derived from the nucleotide inosine monophosphate (IMP), which is the first compound in the pathway to have a completely formed purine ring system.
Inosine monophosphate is synthesized on a pre-existing ribose-phosphate through a complex pathway (as shown in the figure on the right). The source of the carbon and nitrogen atoms of the purine ring, 5 and 4 respectively, come from multiple sources. The amino acid glycine contributes all its carbon (2) and nitrogen (1) atoms, with additional nitrogen atoms from glutamine (2) and aspartic acid (1), and additional carbon atoms from formyl groups (2), which are transferred from the coenzyme tetrahydrofolate as 10-formyltetrahydrofolate, and a carbon atom from bicarbonate (1). Formyl groups build carbon-2 and carbon-8 in the purine ring system, which are the ones acting as bridges between two nitrogen atoms.
GMP
- IMP dehydrogenase converts IMP into XMP
- GMP synthase converts XMP into GMP
- GMP reductase converts GMP back into IMP
AMP
- adenylosuccinate synthase converts IMP to adenylosuccinate
- adenylosuccinate lyase converts adenylosuccinate into AMP
- AMP deaminase converts AMP back into IMP
Degradation
Purines are metabolised by several enzymes:
Guanine
- A nuclease frees the nucleotide
- A nucleotidase creates guanosine
- Purine nucleoside phosphorylase converts guanosine to guanine
- Guanase converts guanine to xanthine
- Xanthine oxidase (a form of xanthine oxidoreductase) catalyzes the oxidation of xanthine to uric acid
Adenine
- A nuclease frees the nucleotide
- A nucleotidase creates adenosine, then adenosine deaminase creates inosine
- Alternatively, AMP deaminase creates inosinic acid, then a nucleotidase creates inosine
- Purine nucleoside phosphorylase acts upon inosine to create hypoxanthine
- Xanthine oxidase catalyzes the biotransformation of hypoxanthine to xanthine
- Xanthine oxidase acts upon xanthine to create uric acid
Salvage
Purines from turnover of nucleic acids (or from food) can also be salvaged and reused in new nucleotides.
- The enzyme adenine phosphoribosyltransferase (APRT) salvages adenine.
- The enzyme hypoxanthine-guanine phosphoribosyltransferase (HGPRT) salvages guanine and hypoxanthine.[2] (Genetic deficiency of HGPRT causes Lesch-Nyhan syndrome.)
Disorders
When a defective gene causes gaps to appear in the metabolic recycling process for purines and pyrimidines, these chemicals are not metabolised properly, and adults or children can suffer from any one of twenty-eight hereditary disorders, possibly some more as yet unknown. Symptoms can include gout, anaemia, epilepsy, delayed development, deafness, compulsive self-biting, kidney failure or stones, or loss of immunity.
Pharmacotherapy
Modulation of purine metabolism has pharmacotherapeutic value.
Purine synthesis inhibitors inhibit the proliferation of cells, especially leukocytes. These inhibitors include azathioprine, an immunosuppressant used in organ transplantation, autoimmune disease such as rheumatoid arthritis or inflammatory bowel disease such as Crohn's disease and ulcerative colitis.
Mycophenolate mofetil is an immunosuppressant drug used to prevent rejection in organ transplantation; it inhibits purine synthesis by blocking inositol monophosphate dehydrogenase. Also Methotrexate indirectly inhibits purine synthesis by blocking the metabolism of folic acid (it is an inhibitor of the Dihydrofolate reductase).
Allopurinol is a drug that inhibits the enzyme xanthine oxidoreductase and, thus, lowers the level of uric acid in the body. This may be useful in the treatment of gout, which is a disease caused by excess uric acid, forming crystals in joints.
See also
References
- ↑ "Substrate specificity of glycinamide ribonucleotide synthetase from chicken liver". The Journal of Biological Chemistry. 271: 8192–5. April 5, 1996. doi:10.1074/jbc.271.14.8192. PMID 8626510.
- ↑ Ansari MY, Equbal A, Dikhit MR, Mansuri R, Rana S, Ali V, Sahoo GC, Das P (Nov 2015). "Establishment of Correlation between In-Silico &In-Vitro Test Analysis against Leishmania HGPRT to inhibitors". International Journal of Biological Macromolecules. 83: 78–96. doi:10.1016/j.ijbiomac.2015.11.051. PMID 26616453.
External links
- The Medical Biochemistry Page
- Purine metabolism - Reference pathway
- PUMPA: Purine Metabolic Patients’ Association